Cis-Allosteric Regulation of HIV-1 Reverse Transcriptase by Integrase
- PMID: 36680070
- PMCID: PMC9864105
- DOI: 10.3390/v15010031
Cis-Allosteric Regulation of HIV-1 Reverse Transcriptase by Integrase
Abstract
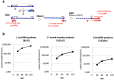
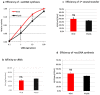
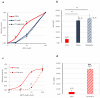
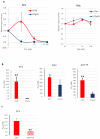
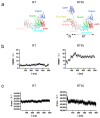
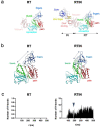
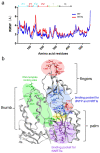
Reverse transcriptase (RT) and integrase (IN) are encoded tandemly in the pol genes of retroviruses. We reported recently that HIV-1 RT and IN need to be supplied as the pol precursor intermediates, in which RT and IN are in fusion form (RTIN) to exert efficient reverse transcription in the context of HIV-1 replication. The mechanism underlying RTIN's effect, however, remains to be elucidated. In this study, we examined the effect of IN fusion on RT during reverse transcription by an in vitro cell-free assay, using recombinant HIV-1 RTIN (rRTIN). We found that, compared to recombinant RT (rRT), rRTIN generated significantly higher cDNAs under physiological concentrations of dNTPs (less than 10 μM), suggesting increased affinity of RTIN to dNTPs. Importantly, the cleavage of RTIN with HIV-1 protease reduced cDNA levels at a low dose of dNTPs. Similarly, sensitivities against RT inhibitors were significantly altered in RTIN form. Finally, analysis of molecular dynamics simulations of RT and RTIN suggested that IN can influence the structural dynamics of the RT active center and the inhibitor binding pockets in cis. Thus, we demonstrated, for the first time, the cis-allosteric regulatory roles of IN in RT structure and enzymatic activity.
Keywords: HIV-1; deoxyribonucleoside triphosphates; integrase; integration; molecular dynamics; pol; protease; reverse transcriptase.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Brown P.O. Integration. In: Coffin J.M., Hughes S.H., Varmus H.E., editors. Retroviruses. Cold Spring Harbor Laboratory Press; Long Island, NY, USA: 1997. pp. 161–203. - PubMed
-
- Tsurutani N., Kubo M., Maeda Y., Ohashi T., Yamamoto N., Kannagi M., Masuda T. Identification of Critical Amino Acid Residues in Human Immunodeficiency Virus Type 1 IN Required for Efficient Proviral DNA Formation at Steps prior to Integration in Dividing and Nondividing Cells. J. Virol. 2000;74:4795–4806. doi: 10.1128/jvi.74.10.4795-4806.2000. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

