The root enrichment of bacteria is consistent across different stress-resistant plant species
- PMID: 36684671
- PMCID: PMC9854377
- DOI: 10.7717/peerj.14683
The root enrichment of bacteria is consistent across different stress-resistant plant species
Abstract
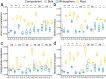
Bacteria, inhabiting around and in plant roots, confer many beneficial traits to promote plant growth and health. The secretion of root exudates modulates the nutritional state of the rhizosphere and root area, further selecting specific bacteria taxa and shaping the bacteria communities. Many studies of the rhizosphere effects have demonstrated that selection by the plant rhizosphere consistently enriches a set of bacteria taxa, and this is conserved across different plant species. Root selection effects are considered to be stronger than the rhizosphere selection effects, yet studies are limited. Here, we focus on the root selection effects across a group of 11 stress-resistant plant species. We found that the root selection consistently reduced the alpha diversity (represented by total number of observed species, Shannon's diversity, and phylogenetic diversity) and altered the structure and composition of bacteria communities. Furthermore, root selection tended to enrich for clusters of bacteria genera including Pantoea, Akkermansia, Blautia, Acinetobacter, Burkholderia-Paraburkholderia, Novosphingobium, Massilia, Pseudomonas, Chryseobacterium, and Stenotrophomonas. Our study offers some basic knowledge for understanding the microbial ecology of the plant root, and suggests that several bacteria genera are of interest for future studies.
Keywords: Abiotic stress; Amplicon sequencing; Bacteria; Microbial diversity; Microbial richness; Microbiota; Pantoea; Plant-microbe interaction; Rhizosphere community; Root.
©2023 Huang et al.
Conflict of interest statement
The authors declare there are no competing interests.
Figures





References
-
- Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R. QIIME allows analysis of high-throughput community sequencing data. Nature Methods. 2010;7:335–336. doi: 10.1038/nmeth.f.303. - DOI - PMC - PubMed
MeSH terms
Associated data
LinkOut - more resources
Full Text Sources

