Single-cell transcriptome sequencing atlas of cassava tuberous root
- PMID: 36684718
- PMCID: PMC9848496
- DOI: 10.3389/fpls.2022.1053669
Single-cell transcriptome sequencing atlas of cassava tuberous root
Abstract
Introduction: Single-cell transcriptome sequencing (ScRNA-seq) has emerged as an effective method for examining cell differentiation and development. In non-model plants, it hasn't been employed very much, especially in sink organs that are abundant in secondary metabolites.
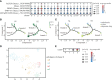
Results: In this study, we sequenced the single-cell transcriptomes at two developmental phases of cassava tuberous roots using the technology known as 10x Genomics (S1, S2). In total, 14,566 cells were grouped into 15 different cell types, primarily based on the marker genes of model plants known to exist. In the pseudotime study, the cell differentiation trajectory was defined, and the difference in gene expression between the two stages on the pseudotime axis was compared. The differentiation process of the vascular tissue and cerebral tissue was identified by the trajectory. We discovered the rare cell type known as the casparian strip via the use of up-regulated genes and pseudotime analysis, and we explained how it differentiates from endodermis. The successful creation of a protoplast isolation technique for organs rich in starch was also described in our study.
Discussion: Together, we created the first high-resolution single-cell transcriptome atlas of cassava tuberous roots, which made significant advancements in our understanding of how these roots differentiate and develop.
Keywords: ScRNA-seq; casparian strip; cassava; differentiation trajectory; tuberous root.
Copyright © 2023 Song, Fan, Shao, Zang, Wang and Min.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



References
LinkOut - more resources
Full Text Sources

