In silico and in vitro studies of potential inhibitors against Dengue viral protein NS5 Methyl Transferase from Ginseng and Notoginseng
- PMID: 36685072
- PMCID: PMC9845645
- DOI: 10.1016/j.jtcme.2022.12.002
In silico and in vitro studies of potential inhibitors against Dengue viral protein NS5 Methyl Transferase from Ginseng and Notoginseng
Abstract
Background and aim: Dengue is a potentially deadly tropical infectious disease transmitted by mosquito vector Aedes aegypti with no antiviral drug available to date. Dengue NS5 protein is crucial for viral replication and is the most conserved among all four Dengue serotypes, making it an attractive drug target. Both Ginseng and Notoginseng extracts and isolates have been shown to be effective against various viral infections yet against Dengue Virus is understudied. We aim to identify potential inhibitors against Dengue NS5 Methyl transferase from small molecular compounds found in Ginseng and Notoginseng.
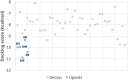
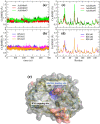
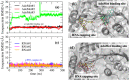
Experimental procedure: A molecular docking model of Dengue NS5 Methyl transferase (MTase) domain was tested with decoys and then used to screen 91 small molecular compounds found in Ginseng and Notoginseng followed by Molecular dynamics simulations and the per-residue free energy decompositions based on molecular mechanics/Poisson-Boltzmann (generalised Born) surface area (MM/PB(GB)SA) calculations of the hit. ADME predictions and drug-likeness analyses were discussed to evaluate the viability of the hit as a drug candidate. To confirm our findings, in vitro studies of antiviral activities against RNA and a E protein synthesis and cell toxicity were carried out.
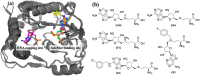
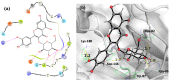
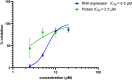
Results and conclusion: The virtual screening resulted in Isoquercitrin as a single hit. Further analyses of the Isoquercitrin-MTase complex show that Isoquercitrin can reside within both of the NS5 Methyl Transferase active sites; the AdoMet binding site and the RNA capping site. The Isoquercitrin is safe for consumption and accessible on multikilogram scale. In vitro studies showed that Isoquercitrin can inhibit Dengue virus by reducing viral RNA and viral protein synthesis with low toxicity to cells (CC50 > 20 μM). Our work provides evidence that Isoquercitrin can serve as an inhibitor of Dengue NS5 protein at the Methyl Transferase domain, further supporting its role as an anti-DENV agent.
Keywords: Antiviral; Dengue; Isoquercitrin; Molecular docking; Molecular dynamics simulations; NS5 methyl transferase.
© 2022 Center for Food and Biomolecules, National Taiwan University. Production and hosting by Elsevier Taiwan LLC.
Conflict of interest statement
Authors declare no conflict of interest.
Figures

References
-
- Wu Q., Dong S., Li X., et al. Effects of COVID-19 non-pharmacological interventions on dengue infection: a systematic review and meta-analysis. Front Cell Infect Microbiol. 2022;12 https://www.frontiersin.org/article/10.3389/fcimb.2022.892508 - DOI - PMC - PubMed
-
- Lindenbach B.D., Thiel H.J., Rice C.M. fifth ed. Lippincott-Raven Publishers; 2007. Fields Virology Chapter 33; Flaviviridae: The Viruses and Their Replication.
LinkOut - more resources
Full Text Sources
Research Materials
