Immunodominant antibody responses directed to SARS-CoV-2 hotspot mutation sites and risk of immune escape
- PMID: 36685521
- PMCID: PMC9849925
- DOI: 10.3389/fimmu.2022.1010105
Immunodominant antibody responses directed to SARS-CoV-2 hotspot mutation sites and risk of immune escape
Abstract
Introduction: Considering the likely need for the development of novel effective vaccines adapted to emerging relevant CoV-2 variants, the increasing knowledge of epitope recognition profile among convalescents and afterwards vaccinated with identification of immunodominant regions may provide important information.
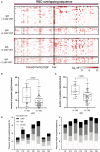
Methods: We used an RBD peptide microarray to identify IgG and IgA binding regions in serum of 71 COVID-19 convalescents and 18 vaccinated individuals.
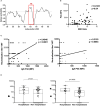
Results: We found a set of immunodominant RBD antibody epitopes, each recognized by more than 30% of the tested cohort, that differ among the two different groups and are within conserved regions among betacoronavirus. Of those, only one peptide, P44 (S415-429), recognized by 68% of convalescents, presented IgG and IgA antibody reactivity that positively correlated with nAb titers, suggesting that this is a relevant RBD region and a potential target of IgG/IgA neutralizing activity.
Discussion: This peptide is localized within the area of contact with ACE-2 and harbors the mutation hotspot site K417 present in gamma (K417T), beta (K417N), and omicron (K417N) variants of concern. The epitope profile of vaccinated individuals differed from convalescents, with a more diverse repertoire of immunodominant peptides, recognized by more than 30% of the cohort. Noteworthy, immunodominant regions of recognition by vaccinated coincide with mutation sites at Omicron BA.1, an important variant emerging after massive vaccination. Together, our data show that immune pressure induced by dominant antibody responses may favor hotspot mutation sites and the selection of variants capable of evading humoral response.
Keywords: RBD; betacoronavirus; immune pressure; linear antibody epitopes; peptide array; sarbecovirus.
Copyright © 2023 Oliveira, Ruiz, Machado, Magawa, Daher, Urbanski, Schmitz, Arcuri, Ferreira, Sasahara, de Medeiros, Júnior, Durigon, Boscardin, Rosa, Schechtman, Nakaya, Cunha-Neto, Gadermaier, Kalil, Coelho and Santos.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures





References
-
- Tai W, He L, Zhang X, Pu J, Voronin D, Jiang S, et al. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol (2020) 17(6). doi: 10.1038/s41423-020-0400-4 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

