A CRISPR way for accelerating cereal crop improvement: Progress and challenges
- PMID: 36685816
- PMCID: PMC9852743
- DOI: 10.3389/fgene.2022.866976
A CRISPR way for accelerating cereal crop improvement: Progress and challenges
Abstract
Humans rely heavily on cereal grains as a key source of nutrients, hence regular improvement of cereal crops is essential for ensuring food security. The current food crisis at the global level is due to the rising population and harsh climatic conditions which prompts scientists to develop smart resilient cereal crops to attain food security. Cereal crop improvement in the past generally depended on imprecise methods like random mutagenesis and conventional genetic recombination which results in high off targeting risks. In this context, we have witnessed the application of targeted mutagenesis using versatile CRISPR-Cas systems for cereal crop improvement in sustainable agriculture. Accelerated crop improvement using molecular breeding methods based on CRISPR-Cas genome editing (GE) is an unprecedented tool for plant biotechnology and agriculture. The last decade has shown the fidelity, accuracy, low levels of off-target effects, and the high efficacy of CRISPR technology to induce targeted mutagenesis for the improvement of cereal crops such as wheat, rice, maize, barley, and millets. Since the genomic databases of these cereal crops are available, several modifications using GE technologies have been performed to attain desirable results. This review provides a brief overview of GE technologies and includes an elaborate account of the mechanisms and applications of CRISPR-Cas editing systems to induce targeted mutagenesis in cereal crops for improving the desired traits. Further, we describe recent developments in CRISPR-Cas-based targeted mutagenesis through base editing and prime editing to develop resilient cereal crop plants, possibly providing new dimensions in the field of cereal crop genome editing.
Keywords: CRISPR/Cas; TALENs; base editing; cereals; crop improvement; food security; genome editing; prime editing.
Copyright © 2023 Basu, Riaz Ahmed, Bhat, Anwar, Ali, Ijaz, Gulzar, Bibi, Tyagi, Nebapure, Goud, Ahanger, Ali and Mushtaq.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

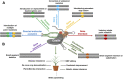
References
Publication types
LinkOut - more resources
Full Text Sources
Miscellaneous

