Evolutionary genomics: Insights from the invasive European starlings
- PMID: 36685843
- PMCID: PMC9845568
- DOI: 10.3389/fgene.2022.1010456
Evolutionary genomics: Insights from the invasive European starlings
Abstract
Two fundamental questions for evolutionary studies are the speed at which evolution occurs, and the way that this evolution may present itself within an organism's genome. Evolutionary studies on invasive populations are poised to tackle some of these pressing questions, including understanding the mechanisms behind rapid adaptation, and how it facilitates population persistence within a novel environment. Investigation of these questions are assisted through recent developments in experimental, sequencing, and analytical protocols; in particular, the growing accessibility of next generation sequencing has enabled a broader range of taxa to be characterised. In this perspective, we discuss recent genetic findings within the invasive European starlings in Australia, and outline some critical next steps within this research system. Further, we use discoveries within this study system to guide discussion of pressing future research directions more generally within the fields of population and evolutionary genetics, including the use of historic specimens, phenotypic data, non-SNP genetic variants (e.g., structural variants), and pan-genomes. In particular, we emphasise the need for exploratory genomics studies across a range of invasive taxa so we can begin understanding broad mechanisms that underpin rapid adaptation in these systems. Understanding how genetic diversity arises and is maintained in a population, and how this contributes to adaptability, requires a deep understanding of how evolution functions at the molecular level, and is of fundamental importance for the future studies and preservation of biodiversity across the globe.
Keywords: Sturnus vulgaris; museum specimens; plasticity; population genetics; rapid adaptation; structural variants.
Copyright © 2023 Stuart, Sherwin, Edwards and Rollins.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

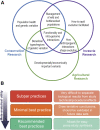
References
-
- Álvarez-Varas R., Heidemeyer M., Riginos C., Benítez H. A., Reséndiz E., Lara-Uc M., et al. (2021). Integrating morphological and genetic data at different spatial scales in a cosmopolitan marine turtle species: Challenges for management and conservation. Zool. J. Linn. Soc. 191, 434–453. 10.1093/zoolinnean/zlaa066 - DOI
-
- Bernstein J. M., Ruane S. (2022). Maximizing molecular data from low-quality fluid-preserved specimens in natural history collections. Front. Ecol. Evol. 10, 893088. 10.3389/fevo.2022.893088 - DOI
-
- Bhattarai G. P., Meyerson L. A., Anderson J., Cummings D., Allen W. J., Cronin J. T. (2017). Biogeography of a plant invasion: Genetic variation and plasticity in latitudinal clines for traits related to herbivory. Ecol. Monogr. 87, 57–75. 10.1002/ecm.1233 - DOI
LinkOut - more resources
Full Text Sources

