Peroxisomal KAT2 (3-ketoacyl-CoA thiolase 2) gene has a key role in gingerol biosynthesis in ginger (Zingiber officinale Rosc.)
- PMID: 36685987
- PMCID: PMC9838548
- DOI: 10.1007/s13562-022-00825-x
Peroxisomal KAT2 (3-ketoacyl-CoA thiolase 2) gene has a key role in gingerol biosynthesis in ginger (Zingiber officinale Rosc.)
Abstract
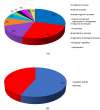
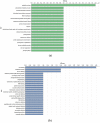
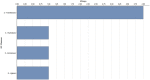
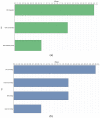
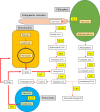
Ginger is an important spice crop with medicinal values and gingerols are the most abundant pungent polyphenols present in ginger, responsible for most of its pharmacological properties. The present study focuses on the molecular mechanism of gingerol biosynthesis in ginger using transcriptome analysis. Suppression Subtractive Hybridization (SSH) was done in leaf and rhizome tissues using high gingerol-producing ginger somaclone B3 as the tester and parent cultivar Maran as the driver and generated high-quality leaf and rhizome Expressed Sequence Tags (ESTs). The Blast2GO annotations of the ESTs revealed the involvement of leaf ESTs in secondary metabolite production, identifying the peroxisomal KAT2 gene (Leaf EST 9) for the high gingerol production in ginger. Rhizome ESTs mostly coded for DNA metabolic processes and differential genes for high gingerol production were not observed in rhizomes. In the qRT-PCR analysis, somaclone B3 had shown high chalcone synthase (CHS: rate-limiting gene in gingerol biosynthetic pathway) activity (0.54 fold) in the leaves of rhizome sprouts. The presence of a high gingerol gene in leaf ESTs and high expression of CHS in leaves presumed that the site of synthesis of gingerols in ginger is the leaves. A modified pathway for gingerol/polyketide backbone formation has been constructed explaining the involvement of KAT gene isoforms KAT2 and KAT5 in gingerol/flavonoid biosynthesis, specifically the KAT2 gene which is otherwise thought to be involved mainly in β-oxidation. The results of the present investigations have the potential of utilizing KAT/thiolase superfamily enzymes for protein/metabolic pathway engineering in ginger for large-scale production of gingerols.
Supplementary information: The online version contains supplementary material available at 10.1007/s13562-022-00825-x.
Keywords: 3-ketoacyl-CoA thiolase/KAT; Expressed Sequence Tags/ESTs; Ginger; Gingerol; Suppression subtractive hybridization; cDNA library.
© The Author(s), under exclusive licence to Society for Plant Biochemistry and Biotechnology 2023, Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Conflict of interest statement
Conflict of interestsThe authors have no competing interests to declare that are relevant to the content of this article.
Figures

Similar articles
-
Transcriptome Analysis Provides Insights into Gingerol Biosynthesis in Ginger (Zingiber officinale).Plant Genome. 2018 Nov;11(3). doi: 10.3835/plantgenome2018.06.0034. Plant Genome. 2018. PMID: 30512040
-
Ginger and turmeric expressed sequence tags identify signature genes for rhizome identity and development and the biosynthesis of curcuminoids, gingerols and terpenoids.BMC Plant Biol. 2013 Feb 15;13:27. doi: 10.1186/1471-2229-13-27. BMC Plant Biol. 2013. PMID: 23410187 Free PMC article.
-
Biosynthesis of curcuminoids and gingerols in turmeric (Curcuma longa) and ginger (Zingiber officinale): identification of curcuminoid synthase and hydroxycinnamoyl-CoA thioesterases.Phytochemistry. 2006 Sep;67(18):2017-29. doi: 10.1016/j.phytochem.2006.06.028. Epub 2006 Aug 7. Phytochemistry. 2006. PMID: 16890967
-
Revisiting the therapeutic potential of gingerols against different pharmacological activities.Naunyn Schmiedebergs Arch Pharmacol. 2023 Apr;396(4):633-647. doi: 10.1007/s00210-022-02372-7. Epub 2022 Dec 31. Naunyn Schmiedebergs Arch Pharmacol. 2023. PMID: 36585999 Free PMC article. Review.
-
Ginger (Zingiber officinale Rosc.) and its bioactive components are potential resources for health beneficial agents.Phytother Res. 2021 Feb;35(2):711-742. doi: 10.1002/ptr.6858. Epub 2020 Sep 20. Phytother Res. 2021. PMID: 32954562
References
-
- Almatroodi SA, Alnuqaydan AM, Babiker AY, Almogbel MA, Khan AA, Rahmani AH. 6-gingerol, a bioactive compound of ginger attenuates renal damage in streptozotocin-induced diabetic rats by regulating the oxidative stress and inflammation. Pharm. 2021;13:317. doi: 10.3390/pharmaceutics13030317. - DOI - PMC - PubMed
-
- Assembly [Internet]. Bethesda (MD): National Library of Medicine (US), National Center for Biotechnology Information; [2021] – Accession No. GCF_018446385.1, Zingiber officinale, chromosome; [cited 2021/05/21]. Available from: https://www.ncbi.nlm.nih.gov/assembly/GCF_018446385.1
-
- Awasthi P, Singh A, Sheikh G, Mahajan V, Gupta AP, Gupta S, Bedi YS, Gandhi SG. Mining and characterization of EST-SSR markers for Zingiber officinale Roscoe with transferability to other species of Zingiberaceae. Physiol Mol Biol Plants. 2017;23(4):925–931. doi: 10.1007/s12298-017-0472-5. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
