Cortical glutamatergic projection neuron types contribute to distinct functional subnetworks
- PMID: 36690901
- PMCID: PMC10571488
- DOI: 10.1038/s41593-022-01244-w
Cortical glutamatergic projection neuron types contribute to distinct functional subnetworks
Abstract
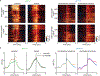
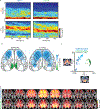
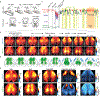
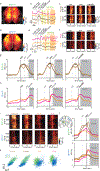
The cellular basis of cerebral cortex functional architecture remains not well understood. A major challenge is to monitor and decipher neural network dynamics across broad cortical areas yet with projection-neuron-type resolution in real time during behavior. Combining genetic targeting and wide-field imaging, we monitored activity dynamics of subcortical-projecting (PTFezf2) and intratelencephalic-projecting (ITPlxnD1) types across dorsal cortex of mice during different brain states and behaviors. ITPlxnD1 and PTFezf2 neurons showed distinct activation patterns during wakeful resting, during spontaneous movements and upon sensory stimulation. Distinct ITPlxnD1 and PTFezf2 subnetworks were dynamically tuned to different sensorimotor components of a naturalistic feeding behavior, and optogenetic inhibition of ITsPlxnD1 and PTsFezf2 in subnetwork nodes disrupted distinct components of this behavior. Lastly, ITPlxnD1 and PTFezf2 projection patterns are consistent with their subnetwork activation patterns. Our results show that, in addition to the concept of columnar organization, dynamic areal and projection-neuron-type specific subnetworks are a key feature of cortical functional architecture linking microcircuit components with global brain networks.
© 2023. The Author(s), under exclusive licence to Springer Nature America, Inc.
Conflict of interest statement
Competing interest
The authors declare no competing interests
Figures









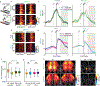


References
-
- Hubel David H & Wiesel Torsten N (New York, NY: Oxford University Press, 2005).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

