Identification of biomarkers related to copper metabolism in patients with pulmonary arterial hypertension
- PMID: 36690956
- PMCID: PMC9868507
- DOI: 10.1186/s12890-023-02326-6
Identification of biomarkers related to copper metabolism in patients with pulmonary arterial hypertension
Abstract
Background: The pathogenesis of pulmonary arterial hypertension (PAH) and associated biomarkers remain to be studied. Copper metabolism is an emerging metabolic research direction in many diseases, but its role in PAH is still unclear.
Methods: PAH-related datasets were downloaded from the Gene Expression Omnibus database, and 2067 copper metabolism-related genes (CMGs) were obtained from the GeneCards database. Differential expression analysis and the Venn algorithm were used to acquire the differentially expressed CMGs (DE-CMGs). DE-CMGs were then used for the coexpression network construction to screen candidate key genes associated with PAH. Furthermore, the predictive performance of the model was verified by receiver operating characteristic (ROC) analysis, and genes with area under the curve (AUC) values greater than 0.8 were selected as diagnostic genes. Then support vector machine, least absolute shrinkage and selection operator regression, and Venn diagrams were applied to detect biomarkers. Moreover, gene set enrichment analysis was performed to explore the function of the biomarkers, and immune-related analyses were utilized to study the infiltration of immune cells. The drug-gene interaction database was used to predict potential therapeutic drugs for PAH using the biomarkers. Biomarkers expression in clinical samples was verified by real-time quantitative PCR.
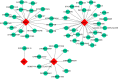
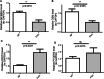
Results: Four biomarkers (DDIT3, NFKBIA, OSM, and PTGER4) were screened. The ROC analysis showed that the 4 biomarkers performed well (AUCs > 0.7). The high expression groups for the 4 biomarkers were enriched in protein activity-related pathways including protein export, spliceosome and proteasome. Furthermore, 8 immune cell types were significantly different between the two groups, including naive B cells, memory B cells, and resting memory CD4 T cells. Afterward, a gene-drug network was constructed. This network illustrated that STREPTOZOCIN, IBUPROFEN, and CELECOXIB were shared by the PTGER4 and DDIT3. Finally, the results of RT-qPCR in clinical samples further confirmed the results of the public database for the expression of NFKBIA and OSM.
Conclusion: In conclusion, four biomarkers (DDIT3, NFKBIA, OSM, and PTGER4) with considerable diagnostic values were identified, and a gene-drug network was further constructed. The results of this study may have significant implications for the development of new diagnostic biomarkers and actionable targets to expand treatment options for PAH patients.
Keywords: Biomarkers; Copper metabolism-related genes; Pulmonary arterial hypertension.
© 2023. The Author(s).
Conflict of interest statement
The authors have no relevant interests of financial or non-financial interests to declared.
Figures







Similar articles
-
Bioinformatics analysis of the immune cell infiltration characteristics and correlation with crucial diagnostic markers in pulmonary arterial hypertension.BMC Pulm Med. 2023 Aug 15;23(1):300. doi: 10.1186/s12890-023-02584-4. BMC Pulm Med. 2023. PMID: 37582718 Free PMC article.
-
Lasso algorithm and support vector machine strategy to screen pulmonary arterial hypertension gene diagnostic markers.Scott Med J. 2023 Feb;68(1):21-31. doi: 10.1177/00369330221132158. Epub 2022 Oct 17. Scott Med J. 2023. PMID: 36253715
-
Analysis of m7G methylation modification patterns and pulmonary vascular immune microenvironment in pulmonary arterial hypertension.Front Immunol. 2022 Dec 5;13:1014509. doi: 10.3389/fimmu.2022.1014509. eCollection 2022. Front Immunol. 2022. PMID: 36544768 Free PMC article.
-
Screening of key biomarkers and immune infiltration in Pulmonary Arterial Hypertension via integrated bioinformatics analysis.Bioengineered. 2021 Dec;12(1):2576-2591. doi: 10.1080/21655979.2021.1936816. Bioengineered. 2021. PMID: 34233597 Free PMC article.
-
Pursuing functional biomarkers in complex disease: Focus on pulmonary arterial hypertension.Am Heart J. 2023 Apr;258:96-113. doi: 10.1016/j.ahj.2022.12.009. Epub 2022 Dec 22. Am Heart J. 2023. PMID: 36565787 Review.
Cited by
-
Inhibition of pulmonary artery smooth muscle cells via the delivery of curcuminoid WZ35 by Cu-based metal organic frameworks.IET Nanobiotechnol. 2023 Jul;17(5):420-424. doi: 10.1049/nbt2.12138. Epub 2023 May 16. IET Nanobiotechnol. 2023. PMID: 37194386 Free PMC article.
-
Emerging trends in cardiovascular diseases: the impact of ferroptosis and cuproptosis on cardiomyocyte death.Mol Cell Biochem. 2025 Jun 23. doi: 10.1007/s11010-025-05340-w. Online ahead of print. Mol Cell Biochem. 2025. PMID: 40549305 Review.
-
Copper homeostasis dysregulation in respiratory diseases: a review of current knowledge.Front Physiol. 2024 May 31;15:1243629. doi: 10.3389/fphys.2024.1243629. eCollection 2024. Front Physiol. 2024. PMID: 38883186 Free PMC article. Review.
-
A hypothesis: Potential contributions of metals to the pathogenesis of pulmonary artery hypertension.Life Sci. 2024 Jan 1;336:122289. doi: 10.1016/j.lfs.2023.122289. Epub 2023 Nov 24. Life Sci. 2024. PMID: 38007143 Free PMC article. Review.
-
KnockTF 2.0: a comprehensive gene expression profile database with knockdown/knockout of transcription (co-)factors in multiple species.Nucleic Acids Res. 2024 Jan 5;52(D1):D183-D193. doi: 10.1093/nar/gkad1016. Nucleic Acids Res. 2024. PMID: 37956336 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

