T7Max transcription system
- PMID: 36691081
- PMCID: PMC9872363
- DOI: 10.1186/s13036-023-00323-1
T7Max transcription system
Abstract
Background: Efficient cell-free protein expression from linear DNA templates has remained a challenge primarily due to template degradation. In addition, the yields of transcription in cell-free systems lag behind transcriptional efficiency of live cells. Most commonly used in vitro translation systems utilize T7 RNA polymerase, which is also the enzyme included in many commercial kits.
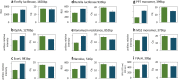
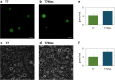
Results: Here we present characterization of a variant of T7 RNA polymerase promoter that acts to significantly increase the yields of gene expression within in vitro systems. We have demonstrated that T7Max increases the yield of translation in many types of commonly used in vitro protein expression systems. We also demonstrated increased protein expression yields from linear templates, allowing the use of T7Max driven expression from linear templates.
Conclusions: The modified promoter, termed T7Max, recruits standard T7 RNA polymerase, so no protein engineering is needed to take advantage of this method. This technique could be used with any T7 RNA polymerase- based in vitro protein expression system.
Keywords: cell-free protein expression; in vitro transcription; in vitro translation; synthetic cells.
© 2023. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





Similar articles
-
Use of T7 RNA polymerase in an optimized Escherichia coli coupled in vitro transcription-translation system. Application in regulatory studies and expression of long transcription units.Eur J Biochem. 1996 Feb 15;236(1):234-9. doi: 10.1111/j.1432-1033.1996.00234.x. Eur J Biochem. 1996. PMID: 8617270
-
In vitro transcription and capping of Gaussia luciferase mRNA followed by HeLa cell transfection.J Vis Exp. 2012 Mar 26;(61):3702. doi: 10.3791/3702. J Vis Exp. 2012. PMID: 22473375 Free PMC article.
-
Preventing nondesired RNA-primed RNA extension catalyzed by T7 RNA polymerase.Eur J Biochem. 2003 Apr;270(7):1458-65. doi: 10.1046/j.1432-1033.2003.03510.x. Eur J Biochem. 2003. PMID: 12654001
-
Bacteriophage T7 transcription system: an enabling tool in synthetic biology.Biotechnol Adv. 2018 Dec;36(8):2129-2137. doi: 10.1016/j.biotechadv.2018.10.001. Epub 2018 Oct 2. Biotechnol Adv. 2018. PMID: 30290194 Review.
-
Towards T7 RNA polymerase (T7RNAP)-based expression system in yeast: challenges and opportunities.Bioengineered. 2022 Jul-Dec;13(7-12):14947-14959. doi: 10.1080/21655979.2023.2180579. Bioengineered. 2022. PMID: 37105766 Free PMC article. Review.
Cited by
-
Innovations, Challenges and Future Directions of T7RNA Polymerase in Microbial Cell Factories.ACS Synth Biol. 2025 May 16;14(5):1381-1396. doi: 10.1021/acssynbio.5c00139. Epub 2025 Apr 10. ACS Synth Biol. 2025. PMID: 40209062 Free PMC article. Review.
-
Multiplexed, image-based pooled screens in primary cells and tissues with PerturbView.Nat Biotechnol. 2025 Jul;43(7):1091-1100. doi: 10.1038/s41587-024-02391-0. Epub 2024 Oct 7. Nat Biotechnol. 2025. PMID: 39375449
-
Parasites, Infections, and Inoculation in Synthetic Minimal Cells.ACS Omega. 2023 Feb 10;8(7):7045-7056. doi: 10.1021/acsomega.2c07911. eCollection 2023 Feb 21. ACS Omega. 2023. PMID: 36844541 Free PMC article.
-
Single cell RNA-seq: a novel tool to unravel virus-host interplay.Virusdisease. 2024 Mar;35(1):41-54. doi: 10.1007/s13337-024-00859-w. Epub 2024 Mar 9. Virusdisease. 2024. PMID: 38817399 Free PMC article. Review.
-
Regulatory Components for Bacterial Cell-Free Systems Engineering.ACS Synth Biol. 2024 Dec 20;13(12):3827-3841. doi: 10.1021/acssynbio.4c00574. Epub 2024 Nov 7. ACS Synth Biol. 2024. PMID: 39509282 Free PMC article. Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

