Rapid nuclear deadenylation of mammalian messenger RNA
- PMID: 36691625
- PMCID: PMC9860345
- DOI: 10.1016/j.isci.2022.105878
Rapid nuclear deadenylation of mammalian messenger RNA
Abstract
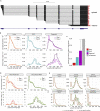
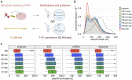
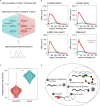
Poly(A) tails protect RNAs from degradation and their deadenylation rates determine RNA stability. Although poly(A) tails are generated in the nucleus, deadenylation of tails has mostly been investigated within the cytoplasm. Here, we combined long-read sequencing with metabolic labeling, splicing inhibition and cell fractionation experiments to quantify, separately, the genesis and trimming of nuclear and cytoplasmic tails in vitro and in vivo. We present evidence for genome-wide, nuclear synthesis of tails longer than 200 nt, which are rapidly shortened after transcription. Our data suggests that rapid deadenylation is a nuclear process, and that different classes of transcripts and even transcript isoforms have distinct nuclear tail lengths. For example, many long-noncoding RNAs retain long poly(A) tails. Modeling deadenylation dynamics predicts nuclear deadenylation about 10 times faster than cytoplasmic deadenylation. In summary, our data suggests that nuclear deadenylation might be a key mechanism for regulating mRNA stability, abundance, and subcellular localization.
Keywords: Cell biology; Molecular mechanism of gene regulation; Molecular physiology.
© 2022 The Authors.
Conflict of interest statement
The authors declare no competing interests. A patent application for FLAM-seq was filed before this work (WO2020069791A1).
Figures


References
-
- Garneau N.L., Wilusz J., Wilusz C.J. The highways and byways of mRNA decay. Nat. Rev. Mol. Cell Biol. 2007;8:113–126. - PubMed
-
- Munroe D., Jacobsen A. Tales of poly(A): a review. Gene. 1990;91:151–158. - PubMed
-
- Weill L., Belloc E., Bava F.A., Méndez R. Translational control by changes in poly(A) tail length: recycling mRNAs. Nat. Struct. Mol. Biol. 2012;19:577–585. - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

