Nodavirus RNA replication crown architecture reveals proto-crown precursor and viral protein A conformational switching
- PMID: 36693094
- PMCID: PMC9945985
- DOI: 10.1073/pnas.2217412120
Nodavirus RNA replication crown architecture reveals proto-crown precursor and viral protein A conformational switching
Abstract
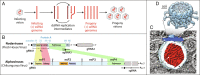
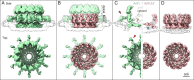
Positive-strand RNA viruses replicate their genomes in virus-induced membrane vesicles, and the resulting RNA replication complexes are a major target for virus control. Nodavirus studies first revealed viral RNA replication proteins forming a 12-fold symmetric "crown" at the vesicle opening to the cytosol, an arrangement recently confirmed to extend to distantly related alphaviruses. Using cryoelectron microscopy (cryo-EM), we show that mature nodavirus crowns comprise two stacked 12-mer rings of multidomain viral RNA replication protein A. Each ring contains an ~19 nm circle of C-proximal polymerase domains, differentiated by strikingly diverged positions of N-proximal RNA capping/membrane binding domains. The lower ring is a "proto-crown" precursor that assembles prior to RNA template recruitment, RNA synthesis, and replication vesicle formation. In this proto-crown, the N-proximal segments interact to form a toroidal central floor, whose 3.1 Å resolution structure reveals many mechanistic details of the RNA capping/membrane binding domains. In the upper ring, cryo-EM fitting indicates that the N-proximal domains extend radially outside the polymerases, forming separated, membrane-binding "legs." The polymerase and N-proximal domains are connected by a long linker accommodating the conformational switch between the two rings and possibly also polymerase movements associated with RNA synthesis and nonsymmetric electron density in the lower center of mature crowns. The results reveal remarkable viral protein multifunctionality, conformational flexibility, and evolutionary plasticity and insights into (+)RNA virus replication and control.
Keywords: RNA replication complex; RNA replication crown; cryo-EM; nodavirus; positive-strand RNA virus.
Conflict of interest statement
The authors declare no competing interest.
Figures







Similar articles
-
Multifunctional Protein A Is the Only Viral Protein Required for Nodavirus RNA Replication Crown Formation.Viruses. 2022 Dec 3;14(12):2711. doi: 10.3390/v14122711. Viruses. 2022. PMID: 36560715 Free PMC article.
-
Subdomain cryo-EM structure of nodaviral replication protein A crown complex provides mechanistic insights into RNA genome replication.Proc Natl Acad Sci U S A. 2020 Aug 4;117(31):18680-18691. doi: 10.1073/pnas.2006165117. Epub 2020 Jul 20. Proc Natl Acad Sci U S A. 2020. PMID: 32690711 Free PMC article.
-
Positive-strand RNA virus genome replication organelles: structure, assembly, control.Trends Genet. 2024 Aug;40(8):681-693. doi: 10.1016/j.tig.2024.04.003. Epub 2024 May 8. Trends Genet. 2024. PMID: 38724328 Review.
-
Cryo-electron microscopy of nodavirus RNA replication organelles illuminates positive-strand RNA virus genome replication.Curr Opin Virol. 2021 Dec;51:74-79. doi: 10.1016/j.coviro.2021.09.008. Epub 2021 Sep 30. Curr Opin Virol. 2021. PMID: 34601307 Free PMC article. Review.
-
Cryo-electron tomography reveals novel features of a viral RNA replication compartment.Elife. 2017 Jun 27;6:e25940. doi: 10.7554/eLife.25940. Elife. 2017. PMID: 28653620 Free PMC article.
Cited by
-
Molecular architecture of coronavirus double-membrane vesicle pore complex.Nature. 2024 Sep;633(8028):224-231. doi: 10.1038/s41586-024-07817-y. Epub 2024 Aug 14. Nature. 2024. PMID: 39143215 Free PMC article.
-
A coronaviral pore-replicase complex links RNA synthesis and export from double-membrane vesicles.Sci Adv. 2024 Nov 8;10(45):eadq9580. doi: 10.1126/sciadv.adq9580. Epub 2024 Nov 8. Sci Adv. 2024. PMID: 39514670 Free PMC article.
-
SARS-CoV-2 nsp3 and nsp4 are minimal constituents of a pore spanning replication organelle.Nat Commun. 2023 Nov 30;14(1):7894. doi: 10.1038/s41467-023-43666-5. Nat Commun. 2023. PMID: 38036567 Free PMC article.
-
Hepatitis E virus RNA replication polyprotein: taking structural biology seriously.Front Microbiol. 2023 Aug 4;14:1254741. doi: 10.3389/fmicb.2023.1254741. eCollection 2023. Front Microbiol. 2023. PMID: 37601361 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

