Protein degradation: expanding the toolbox to restrain cancer drug resistance
- PMID: 36694209
- PMCID: PMC9872387
- DOI: 10.1186/s13045-023-01398-5
Protein degradation: expanding the toolbox to restrain cancer drug resistance
Abstract
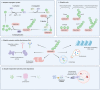
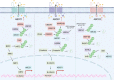
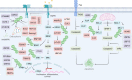
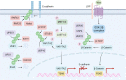
Despite significant progress in clinical management, drug resistance remains a major obstacle. Recent research based on protein degradation to restrain drug resistance has attracted wide attention, and several therapeutic strategies such as inhibition of proteasome with bortezomib and proteolysis-targeting chimeric have been developed. Compared with intervention at the transcriptional level, targeting the degradation process seems to be a more rapid and direct strategy. Proteasomal proteolysis and lysosomal proteolysis are the most critical quality control systems responsible for the degradation of proteins or organelles. Although proteasomal and lysosomal inhibitors (e.g., bortezomib and chloroquine) have achieved certain improvements in some clinical application scenarios, their routine application in practice is still a long way off, which is due to the lack of precise targeting capabilities and inevitable side effects. In-depth studies on the regulatory mechanism of critical protein degradation regulators, including E3 ubiquitin ligases, deubiquitylating enzymes (DUBs), and chaperones, are expected to provide precise clues for developing targeting strategies and reducing side effects. Here, we discuss the underlying mechanisms of protein degradation in regulating drug efflux, drug metabolism, DNA repair, drug target alteration, downstream bypass signaling, sustaining of stemness, and tumor microenvironment remodeling to delineate the functional roles of protein degradation in drug resistance. We also highlight specific E3 ligases, DUBs, and chaperones, discussing possible strategies modulating protein degradation to target cancer drug resistance. A systematic summary of the molecular basis by which protein degradation regulates tumor drug resistance will help facilitate the development of appropriate clinical strategies.
Keywords: Chaperone-mediated autophagy; DUBs; Drug resistance; E3 ligase; Protein degradation.
© 2023. The Author(s).
Conflict of interest statement
The authors declared no potential competing interests.
Figures




Similar articles
-
Perspectives on the development of first-in-class protein degraders.Future Med Chem. 2021 Jul;13(14):1203-1226. doi: 10.4155/fmc-2021-0033. Epub 2021 May 21. Future Med Chem. 2021. PMID: 34015962 Review.
-
E3-ligase knock down revealed differential titin degradation by autopagy and the ubiquitin proteasome system.Sci Rep. 2021 Oct 26;11(1):21134. doi: 10.1038/s41598-021-00618-7. Sci Rep. 2021. PMID: 34702928 Free PMC article.
-
CHIP: a co-chaperone for degradation by the proteasome.Subcell Biochem. 2015;78:219-42. doi: 10.1007/978-3-319-11731-7_11. Subcell Biochem. 2015. PMID: 25487024 Review.
-
It's all about talking: two-way communication between proteasomal and lysosomal degradation pathways via ubiquitin.Am J Physiol Cell Physiol. 2016 Aug 1;311(2):C166-78. doi: 10.1152/ajpcell.00074.2016. Epub 2016 May 25. Am J Physiol Cell Physiol. 2016. PMID: 27225656 Free PMC article.
-
CHIP: A Co-chaperone for Degradation by the Proteasome and Lysosome.Subcell Biochem. 2023;101:351-387. doi: 10.1007/978-3-031-14740-1_12. Subcell Biochem. 2023. PMID: 36520313
Cited by
-
LAMP2A regulates cisplatin resistance in colorectal cancer through mediating autophagy.J Cancer Res Clin Oncol. 2024 May 8;150(5):242. doi: 10.1007/s00432-024-05775-6. J Cancer Res Clin Oncol. 2024. PMID: 38717639 Free PMC article.
-
Crosstalk between O-GlcNAcylation and ubiquitination: a novel strategy for overcoming cancer therapeutic resistance.Exp Hematol Oncol. 2024 Nov 1;13(1):107. doi: 10.1186/s40164-024-00569-5. Exp Hematol Oncol. 2024. PMID: 39487556 Free PMC article. Review.
-
A region-confined PROTAC nanoplatform for spatiotemporally tunable protein degradation and enhanced cancer therapy.Nat Commun. 2024 Aug 4;15(1):6608. doi: 10.1038/s41467-024-50735-w. Nat Commun. 2024. PMID: 39098906 Free PMC article.
-
Cetuximab chemotherapy resistance: Insight into the homeostatic evolution of head and neck cancer (Review).Oncol Rep. 2024 Jun;51(6):80. doi: 10.3892/or.2024.8739. Epub 2024 Apr 19. Oncol Rep. 2024. PMID: 38639184 Free PMC article. Review.
-
A robust multiplex-DIA workflow profiles protein turnover regulations associated with cisplatin resistance and aneuploidy.Nat Commun. 2025 May 30;16(1):5034. doi: 10.1038/s41467-025-60319-x. Nat Commun. 2025. PMID: 40447611 Free PMC article.
References
-
- Dallavalle S, Dobričić V, Lazzarato L, Gazzano E, Machuqueiro M, Pajeva I, Tsakovska I, Zidar N, Fruttero R. Improvement of conventional anti-cancer drugs as new tools against multidrug resistant tumors. Drug Resist Updat Rev Comment Antimicrob Anticancer Chemother. 2020;50:100682. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

