Nanoscopic investigation of C9orf72 poly-GA oligomers on nuclear membrane disruption by a photoinducible platform
- PMID: 36697556
- PMCID: PMC9814621
- DOI: 10.1038/s42004-021-00547-6
Nanoscopic investigation of C9orf72 poly-GA oligomers on nuclear membrane disruption by a photoinducible platform
Abstract
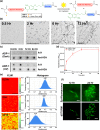
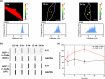
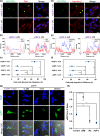
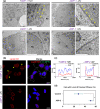
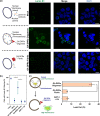
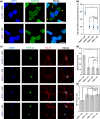
Glycine-alanine dipeptide repeats (GA DPRs) translated from the mutated C9orf72 gene have recently been correlated with amyotrophic lateral sclerosis (ALS). While GA DPRs aggregates have been suggested as amyloid, the biophysical features and cytotoxicity of GA DPRs oligomers has not been explored due to its unstable nature. In this study, we develop a photoinducible platform based on methoxynitrobenzene chemistry to enrich GA DPRs that allows monitoring the oligomerization process of GA DPRs in cells. By applying advanced microscopies, we examined the GA DPRs oligomerization process nanoscopically in a time-dependent manner. We provided direct evidences to demonstrate GA DPRs oligomers rather than nanofibrils disrupt nuclear membrane. Moreover, we found GA DPRs hamper nucleocytoplasmic transport in cells and cause cytosolic retention of TAR DNA-binding protein 43 in cortical neurons. Our results highlight the toxicity of GA DPRs oligomers, which is a key step toward elucidating the pathological roles of C9orf72 DPRs.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Poly-glycine-alanine exacerbates C9orf72 repeat expansion-mediated DNA damage via sequestration of phosphorylated ATM and loss of nuclear hnRNPA3.Acta Neuropathol. 2020 Jan;139(1):99-118. doi: 10.1007/s00401-019-02082-0. Epub 2019 Oct 23. Acta Neuropathol. 2020. PMID: 31642962 Free PMC article.
-
The Glycine-Alanine Dipeptide Repeat from C9orf72 Hexanucleotide Expansions Forms Toxic Amyloids Possessing Cell-to-Cell Transmission Properties.J Biol Chem. 2016 Mar 4;291(10):4903-11. doi: 10.1074/jbc.M115.694273. Epub 2016 Jan 14. J Biol Chem. 2016. PMID: 26769963 Free PMC article.
-
C9orf72 Dipeptide Repeats Cause Selective Neurodegeneration and Cell-Autonomous Excitotoxicity in Drosophila Glutamatergic Neurons.J Neurosci. 2018 Aug 29;38(35):7741-7752. doi: 10.1523/JNEUROSCI.0908-18.2018. Epub 2018 Jul 23. J Neurosci. 2018. PMID: 30037833 Free PMC article.
-
The Role of Dipeptide Repeats in C9ORF72-Related ALS-FTD.Front Mol Neurosci. 2017 Feb 13;10:35. doi: 10.3389/fnmol.2017.00035. eCollection 2017. Front Mol Neurosci. 2017. PMID: 28243191 Free PMC article. Review.
-
C9orf72 ALS-FTD: recent evidence for dysregulation of the autophagy-lysosome pathway at multiple levels.Autophagy. 2021 Nov;17(11):3306-3322. doi: 10.1080/15548627.2021.1872189. Epub 2021 Feb 26. Autophagy. 2021. PMID: 33632058 Free PMC article. Review.
Cited by
-
LINC complex alterations are a key feature of sporadic and familial ALS/FTD.Acta Neuropathol Commun. 2024 Apr 25;12(1):69. doi: 10.1186/s40478-024-01778-z. Acta Neuropathol Commun. 2024. PMID: 38664831 Free PMC article.
-
Glycine-Alanine Dipeptide Repeat Protein from C9-ALS Interacts with Sulfide Quinone Oxidoreductase (SQOR) to Induce the Activity of the NLRP3 Inflammasome in HMC3 Microglia: Irisflorentin Reverses This Interaction.Antioxidants (Basel). 2023 Oct 23;12(10):1896. doi: 10.3390/antiox12101896. Antioxidants (Basel). 2023. PMID: 37891975 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

