Diosgenin enhances liposome-enabled nucleic acid delivery and CRISPR/Cas9-mediated gene editing by modulating endocytic pathways
- PMID: 36698628
- PMCID: PMC9868636
- DOI: 10.3389/fbioe.2022.1031049
Diosgenin enhances liposome-enabled nucleic acid delivery and CRISPR/Cas9-mediated gene editing by modulating endocytic pathways
Abstract
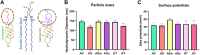
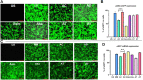
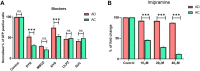
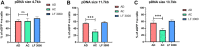
The CRISPR/Cas9 system holds great promise in treating genetic diseases, owing to its safe and precise genome editing. However, the major challenges to implementing the technology in clinics lie in transiently limiting the expression of genome editing factors and achieving therapeutically relevant frequencies with fidelity. Recent findings revealed that non-viral vectors could be a potential alternative delivery system to overcome these limitations. In our previous research, we demonstrated that liposomal formulations with amide linker-based cationic lipids and cholesterol were found to be effective in delivering a variety of nucleic acids. In the current study, we screened steroidal sapogenins as an alternative co-lipid to cholesterol in cationic liposomal formulations and found that liposomes with diosgenin (AD, Amide lipid: Diosgenin) further improved nucleic acid delivery efficacy, in particular, delivering Cas9 pDNA and mRNA for efficient genome editing at multiple loci, including AAVS1 and HBB, when compared to amide cholesterol. Mechanistic insights into the endocytosis of lipoplexes revealed that diosgenin facilitated the lipoplexes' cholesterol-independent and clathrin-mediated endocytosis, which in turn leads to increased intracellular delivery. Our study identifies diosgenin-doped liposomes as an efficient tool to deliver CRISPR/Cas9 system.
Keywords: CRISPR/Cas9; cationic lipid; diosgenin; genome editing; transfections.
Copyright © 2023 Lohchania, Christopher, Arjunan, Mahalingam, Kathirvelu, Prasannan, Venkatesan, Taneja, KM, Thangavel and Marepally.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
-
- Dharmalingam P., Rachamalla H. K. R., Lohchania B., Bandlamudi B., Thangavel S., Murugesan M. K., et al. (2017). Green transfection: Cationic lipid nanocarrier system derivatized from vegetable fat, palmstearin enhances nucleic acid transfections. ACS Omega 2, 7892–7903. 10.1021/acsomega.7b00935 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

