Res-Dom: predicting protein domain boundary from sequence using deep residual network and Bi-LSTM
- PMID: 36699417
- PMCID: PMC9710680
- DOI: 10.1093/bioadv/vbac060
Res-Dom: predicting protein domain boundary from sequence using deep residual network and Bi-LSTM
Abstract
Motivation: Protein domains are the basic units of proteins that can fold, function and evolve independently. Protein domain boundary partition plays an important role in protein structure prediction, understanding their biological functions, annotating their evolutionary mechanisms and protein design. Although there are many methods that have been developed to predict domain boundaries from protein sequence over the past two decades, there is still much room for improvement.
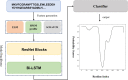
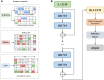
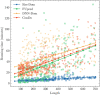
Results: In this article, a novel domain boundary prediction tool called Res-Dom was developed, which is based on a deep residual network, bidirectional long short-term memory (Bi-LSTM) and transfer learning. We used deep residual neural networks to extract higher-order residue-related information. In addition, we also used a pre-trained protein language model called ESM to extract sequence embedded features, which can summarize sequence context information more abundantly. To improve the global representation of these deep residual networks, a Bi-LSTM network was also designed to consider long-range interactions between residues. Res-Dom was then tested on an independent test set including 342 proteins and generated correct single-domain and multi-domain classifications with a Matthew's correlation coefficient of 0.668, which was 17.6% higher than the second-best compared method. For domain boundaries, the normalized domain overlapping score of Res-Dom was 0.849, which was 5% higher than the second-best compared method. Furthermore, Res-Dom required significantly less time than most of the recently developed state-of-the-art domain prediction methods.
Availability and implementation: All source code, datasets and model are available at http://isyslab.info/Res-Dom/.
© The Author(s) 2022. Published by Oxford University Press.
Figures

Similar articles
-
DNN-Dom: predicting protein domain boundary from sequence alone by deep neural network.Bioinformatics. 2019 Dec 15;35(24):5128-5136. doi: 10.1093/bioinformatics/btz464. Bioinformatics. 2019. PMID: 31197306
-
FUpred: detecting protein domains through deep-learning-based contact map prediction.Bioinformatics. 2020 Jun 1;36(12):3749-3757. doi: 10.1093/bioinformatics/btaa217. Bioinformatics. 2020. PMID: 32227201 Free PMC article.
-
Accurate prediction of protein contact maps by coupling residual two-dimensional bidirectional long short-term memory with convolutional neural networks.Bioinformatics. 2018 Dec 1;34(23):4039-4045. doi: 10.1093/bioinformatics/bty481. Bioinformatics. 2018. PMID: 29931279
-
Accurate De Novo Prediction of Protein Contact Map by Ultra-Deep Learning Model.PLoS Comput Biol. 2017 Jan 5;13(1):e1005324. doi: 10.1371/journal.pcbi.1005324. eCollection 2017 Jan. PLoS Comput Biol. 2017. PMID: 28056090 Free PMC article.
-
Protein domain identification methods and online resources.Comput Struct Biotechnol J. 2021 Feb 2;19:1145-1153. doi: 10.1016/j.csbj.2021.01.041. eCollection 2021. Comput Struct Biotechnol J. 2021. PMID: 33680357 Free PMC article. Review.
Cited by
-
Pre-trained protein language model sheds new light on the prediction of Arabidopsis protein-protein interactions.Plant Methods. 2023 Dec 7;19(1):141. doi: 10.1186/s13007-023-01119-6. Plant Methods. 2023. PMID: 38062445 Free PMC article.
-
DeepNeuropePred: A robust and universal tool to predict cleavage sites from neuropeptide precursors by protein language model.Comput Struct Biotechnol J. 2023 Dec 5;23:309-315. doi: 10.1016/j.csbj.2023.12.004. eCollection 2024 Dec. Comput Struct Biotechnol J. 2023. PMID: 38179071 Free PMC article.
References
-
- Alexandrov N., Shindyalov I. (2003) PDP: protein domain parser. Bioinformatics, 19, 429–430. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

