Structural insights into 3Fe-4S ferredoxins diversity in M. tuberculosis highlighted by a first redox complex with P450
- PMID: 36699703
- PMCID: PMC9868604
- DOI: 10.3389/fmolb.2022.1100032
Structural insights into 3Fe-4S ferredoxins diversity in M. tuberculosis highlighted by a first redox complex with P450
Abstract
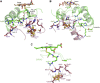
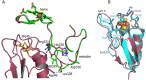
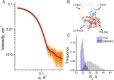
Ferredoxins are small iron-sulfur proteins and key players in essential metabolic pathways. Among all types, 3Fe-4S ferredoxins are less studied mostly due to anaerobic requirements. Their complexes with cytochrome P450 redox partners have not been structurally characterized. In the present work, we solved the structures of both 3Fe-4S ferredoxins from M. tuberculosis-Fdx alone and the fusion FdxE-CYP143. Our SPR analysis demonstrated a high-affinity binding of FdxE to CYP143. According to SAXS data, the same complex is present in solution. The structure reveals extended multipoint interactions and the shape/charge complementarity of redox partners. Furthermore, FdxE binding induced conformational changes in CYP143 as evident from the solved CYP143 structure alone. The comparison of FdxE-CYP143 and modeled Fdx-CYP51 complexes further revealed the specificity of ferredoxins. Our results illuminate the diversity of electron transfer complexes for the production of different secondary metabolites.
Keywords: 3Fe–4S ferredoxins; crystal structure; cytochrome P450; protein–protein interactions; redox complex.
Copyright © 2023 Gilep, Varaksa, Bukhdruker, Kavaleuski, Ryzhykau, Smolskaya, Sushko, Tsumoto, Grabovec, Kapranov, Okhrimenko, Marin, Shevtsov, Mishin, Kovalev, Kuklin, Gordeliy, Kaluzhskiy, Gnedenko, Yablokov, Ivanov, Borshchevskiy and Strushkevich.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
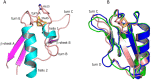



Similar articles
-
Biophysical characterization of the sterol demethylase P450 from Mycobacterium tuberculosis, its cognate ferredoxin, and their interactions.Biochemistry. 2006 Jul 11;45(27):8427-43. doi: 10.1021/bi0601609. Biochemistry. 2006. PMID: 16819841
-
The structure of a novel electron-transfer ferredoxin from Rhodopseudomonas palustris HaA2 which contains a histidine residue in its iron-sulfur cluster-binding motif.Acta Crystallogr D Biol Crystallogr. 2014 May;70(Pt 5):1453-64. doi: 10.1107/S139900471400474X. Epub 2014 Apr 30. Acta Crystallogr D Biol Crystallogr. 2014. PMID: 24816113
-
Oxidized and reduced Azotobacter vinelandii ferredoxin I at 1.4 A resolution: conformational change of surface residues without significant change in the [3Fe-4S]+/0 cluster.Biochemistry. 1999 Jun 29;38(26):8228-39. doi: 10.1021/bi983008i. Biochemistry. 1999. PMID: 10387068
-
An overview of the factors playing a role in cytochrome P450 monooxygenase and ferredoxin interactions.Biophys Rev. 2020 Oct;12(5):1217-1222. doi: 10.1007/s12551-020-00749-7. Epub 2020 Sep 3. Biophys Rev. 2020. PMID: 32885385 Free PMC article. Review.
-
Structure and electrochemistry of proteins harboring iron-sulfur clusters of different nuclearities. Part III. [4Fe-4S], [3Fe-4S] and [2Fe-2S] iron-sulfur proteins.J Struct Biol. 2018 Jun;202(3):264-274. doi: 10.1016/j.jsb.2018.03.008. Epub 2018 Mar 27. J Struct Biol. 2018. PMID: 29601957 Review.
Cited by
-
Antimycobacterial Activities of Hydroxamic Acids and Their Iron(II/III), Nickel(II), Copper(II) and Zinc(II) Complexes.Microorganisms. 2023 Oct 23;11(10):2611. doi: 10.3390/microorganisms11102611. Microorganisms. 2023. PMID: 37894269 Free PMC article.
-
Analysis of the Setomimycin Biosynthetic Gene Cluster from Streptomyces nojiriensis JCM3382 and Evaluation of Its α-Glucosidase Inhibitory Activity Using Molecular Docking and Molecular Dynamics Simulations.Int J Mol Sci. 2024 Oct 6;25(19):10758. doi: 10.3390/ijms251910758. Int J Mol Sci. 2024. PMID: 39409089 Free PMC article.
-
Ferredoxins: Functions, Evolution, Potential Applications, and Challenges of Subtype Classification.Curr Issues Mol Biol. 2024 Sep 1;46(9):9659-9673. doi: 10.3390/cimb46090574. Curr Issues Mol Biol. 2024. PMID: 39329926 Free PMC article. Review.
-
An iron-sulfur cluster as a new metal center in a flavodiiron protein.Protein Sci. 2025 Jul;34(7):e70204. doi: 10.1002/pro.70204. Protein Sci. 2025. PMID: 40545700
References
LinkOut - more resources
Full Text Sources

