Integrative analysis of microRNAs and mRNAs reveals the regulatory networks of triterpenoid saponin metabolism in Soapberry (Sapindus mukorossi Gaertn.)
- PMID: 36699854
- PMCID: PMC9869041
- DOI: 10.3389/fpls.2022.1037784
Integrative analysis of microRNAs and mRNAs reveals the regulatory networks of triterpenoid saponin metabolism in Soapberry (Sapindus mukorossi Gaertn.)
Abstract
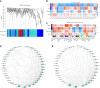
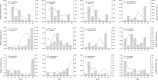
Triterpenoid saponin are important secondary metabolites and bioactive constituents of soapberry (Sapindus mukorossi Gaertn.) and are widely used in medicine and toiletry products. However, little is known about the roles of miRNAs in the regulation of triterpenoid saponin biosynthesis in soapberry. In this study, a total of 3036 miRNAs were identified, of which 1372 miRNAs were differentially expressed at different stages of pericarp development. Important KEGG pathways, such as terpenoid backbone biosynthesis, sesquiterpenoid and triterpenoid biosynthesis, and basal transcription factors were highlighted, as well the roles of some key miRNAs, such as ath-miR5021, han-miR3630-3p, and ppe-miR858, which may play important roles in regulating triterpenoid saponin biosynthesis. In addition, 58 miRNAs might participate in saponin biosynthesis pathways by predicting the targets of those miRNAs to 53 saponin biosynthesis structural genes. And 75 miRNAs were identified to potentially play vital role in saponin accumulation by targeting transcript factor genes, bHLH, bZIP, ERF, MYB, and WRKY, respectively, which are candidate regulatory genes in the pathway of saponin biosynthesis. The results of weighted gene coexpression network analysis (WGCNA) suggested that two saponin-specific miRNA modules and 10 hub miRNAs may participate in saponin biosynthesis. Furthermore, multiple miRNA-mRNA regulatory networks potentially involved in saponin biosynthesis were generated, e.g., ath-miR5021-SmIDI2/SmGPS5/SmbAS1/SmCYP71D-3/SmUGT74G-2, han-miR3630-3p-SmCYP71A-14/SmbHLH54/SmMYB135/SmWRKY32, and ppe-miR858-SmMYB5/SmMYB32. qRT-PCR analysis validated the expression patterns of nine miRNAs and 12 corresponding target genes. This study represents the first comprehensive analysis of miRNAs in soapberry and lays the foundation for further understanding of miRNA-based regulation in triterpenoid saponin biosynthesis.
Keywords: Sapindus mukorossi; biosynthesis; coexpression network; miRNA; triterpenoid saponin.
Copyright © 2023 Xu, Liu, Ji, Zhao, Zhao, Wang, Wang, Gao, Hao, Gao, Gao, Weng, Jia and Chen.
Conflict of interest statement
XJ and XHW were employed by Liming Jia. Author XJ was employed by Hangzhou KaiTai Biotechnology Co., Ltd. Author XHW was employed by Yuanhua Forestry Biological Technology Co., Ltd. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Metabolome and Transcriptome Analysis Reveals the Transcriptional Regulatory Mechanism of Triterpenoid Saponin Biosynthesis in Soapberry (Sapindus mukorossi Gaertn.).J Agric Food Chem. 2022 Jun 15;70(23):7095-7109. doi: 10.1021/acs.jafc.2c01672. Epub 2022 May 31. J Agric Food Chem. 2022. PMID: 35638867
-
Identification and analysis of UGT genes associated with triterpenoid saponin in soapberry (Sapindus mukorossi Gaertn.).BMC Plant Biol. 2024 Jun 21;24(1):588. doi: 10.1186/s12870-024-05281-4. BMC Plant Biol. 2024. PMID: 38902602 Free PMC article.
-
Metabolomics analysis of the soapberry (Sapindus mukorossi Gaertn.) pericarp during fruit development and ripening based on UHPLC-HRMS.Sci Rep. 2021 Jun 2;11(1):11657. doi: 10.1038/s41598-021-91143-0. Sci Rep. 2021. PMID: 34079016 Free PMC article.
-
Triterpenoid Saponins from Washnut (Sapindus mukorossi Gaertn.)-A Source of Natural Surfactants and Other Active Components.Plants (Basel). 2022 Sep 9;11(18):2355. doi: 10.3390/plants11182355. Plants (Basel). 2022. PMID: 36145756 Free PMC article. Review.
-
[The glycosyltransferases involved in triterpenoid saponin biosynthesis: a review].Sheng Wu Gong Cheng Xue Bao. 2022 Mar 25;38(3):1004-1024. doi: 10.13345/j.cjb.210587. Sheng Wu Gong Cheng Xue Bao. 2022. PMID: 35355470 Review. Chinese.
Cited by
-
Effect of nitrogen, phosphorus and potassium fertilization management on soil properties and leaf traits and yield of Sapindus mukorossi.Front Plant Sci. 2024 Mar 11;15:1300683. doi: 10.3389/fpls.2024.1300683. eCollection 2024. Front Plant Sci. 2024. PMID: 38529062 Free PMC article.
-
Integrated miRNAome, transcriptome, and degradome analyses reveal the role of miRNA-mRNA modules in the biosynthesis of oridonin in Isodon rubescens.Front Plant Sci. 2025 Jun 18;16:1566354. doi: 10.3389/fpls.2025.1566354. eCollection 2025. Front Plant Sci. 2025. PMID: 40606481 Free PMC article.
-
Fertilization Induced Soil Microbial Shifts Show Minor Effects on Sapindus mukorossi Yield.Microorganisms. 2025 Jan 15;13(1):173. doi: 10.3390/microorganisms13010173. Microorganisms. 2025. PMID: 39858941 Free PMC article.
References
LinkOut - more resources
Full Text Sources

