Applications of generative adversarial networks in neuroimaging and clinical neuroscience
- PMID: 36702211
- PMCID: PMC9992336
- DOI: 10.1016/j.neuroimage.2023.119898
Applications of generative adversarial networks in neuroimaging and clinical neuroscience
Abstract
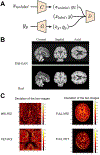
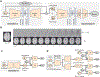
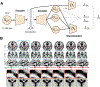
Generative adversarial networks (GANs) are one powerful type of deep learning models that have been successfully utilized in numerous fields. They belong to the broader family of generative methods, which learn to generate realistic data with a probabilistic model by learning distributions from real samples. In the clinical context, GANs have shown enhanced capabilities in capturing spatially complex, nonlinear, and potentially subtle disease effects compared to traditional generative methods. This review critically appraises the existing literature on the applications of GANs in imaging studies of various neurological conditions, including Alzheimer's disease, brain tumors, brain aging, and multiple sclerosis. We provide an intuitive explanation of various GAN methods for each application and further discuss the main challenges, open questions, and promising future directions of leveraging GANs in neuroimaging. We aim to bridge the gap between advanced deep learning methods and neurology research by highlighting how GANs can be leveraged to support clinical decision making and contribute to a better understanding of the structural and functional patterns of brain diseases.
Keywords: GAN; Generative adversarial network; Neuroimaging; Pathology; Review.
Copyright © 2023 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest The authors report no competing interest.
Figures
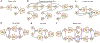


Similar articles
-
Generative Adversarial Network (GAN) for Simulating Electroencephalography.Brain Topogr. 2023 Sep;36(5):661-670. doi: 10.1007/s10548-023-00986-5. Epub 2023 Jul 6. Brain Topogr. 2023. PMID: 37410276
-
Generative Adversarial Networks in Digital Histopathology: Current Applications, Limitations, Ethical Considerations, and Future Directions.Mod Pathol. 2024 Jan;37(1):100369. doi: 10.1016/j.modpat.2023.100369. Epub 2023 Oct 27. Mod Pathol. 2024. PMID: 37890670 Review.
-
A Future Picture: A Review of Current Generative Adversarial Neural Networks in Vitreoretinal Pathologies and Their Future Potentials.Biomedicines. 2025 Jan 24;13(2):284. doi: 10.3390/biomedicines13020284. Biomedicines. 2025. PMID: 40002698 Free PMC article. Review.
-
A review of the applications of generative adversarial networks to structural and functional MRI based diagnostic classification of brain disorders.Front Neurosci. 2024 Apr 15;18:1333712. doi: 10.3389/fnins.2024.1333712. eCollection 2024. Front Neurosci. 2024. PMID: 38686334 Free PMC article. Review.
-
Deepfakes in Ophthalmology: Applications and Realism of Synthetic Retinal Images from Generative Adversarial Networks.Ophthalmol Sci. 2021 Nov 16;1(4):100079. doi: 10.1016/j.xops.2021.100079. eCollection 2021 Dec. Ophthalmol Sci. 2021. PMID: 36246951 Free PMC article.
Cited by
-
From knowledge generation to knowledge verification: examining the biomedical generative capabilities of ChatGPT.iScience. 2025 Apr 21;28(6):112492. doi: 10.1016/j.isci.2025.112492. eCollection 2025 Jun 20. iScience. 2025. PMID: 40487456 Free PMC article.
-
Advancements in Artificial Intelligence for Fetal Neurosonography: A Comprehensive Review.J Clin Med. 2024 Sep 22;13(18):5626. doi: 10.3390/jcm13185626. J Clin Med. 2024. PMID: 39337113 Free PMC article. Review.
-
Uncover This Tech Term: Generative Adversarial Networks.Korean J Radiol. 2024 May;25(5):493-498. doi: 10.3348/kjr.2023.1306. Epub 2024 Apr 11. Korean J Radiol. 2024. PMID: 38627875 Free PMC article. No abstract available.
-
Reversibility of Tumor-like Lesions: Meticulous Diagnosis for Treatable Brain Diseases.Neurol Clin Pract. 2023 Oct;13(5):e200183. doi: 10.1212/CPJ.0000000000200183. Epub 2023 Aug 30. Neurol Clin Pract. 2023. PMID: 37664133 Free PMC article. No abstract available.
-
Advances in Neuroimaging and Deep Learning for Emotion Detection: A Systematic Review of Cognitive Neuroscience and Algorithmic Innovations.Diagnostics (Basel). 2025 Feb 13;15(4):456. doi: 10.3390/diagnostics15040456. Diagnostics (Basel). 2025. PMID: 40002607 Free PMC article. Review.
References
-
- Arjovsky M, Chintala S, Bottou L, 2017. Wasserstein generative adversarial networks. In: Proceedings of the 34th International Conference on Machine Learning PMLR.
-
- Bakas S, Reyes M, Jakab A, Bauer S, Rempfler M, Crimi A, Jambawalikar SR (2018). Identifying the best machine learning algorithms for brain tumor segmentation, progression assessment, and overall survival prediction in the BRATS challenge. arXiv preprint arXiv: 1811.02629
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

