Hotspot of de novo telomere addition stabilizes linear amplicons in yeast grown in sulfate-limiting conditions
- PMID: 36702776
- PMCID: PMC10213492
- DOI: 10.1093/genetics/iyad010
Hotspot of de novo telomere addition stabilizes linear amplicons in yeast grown in sulfate-limiting conditions
Abstract
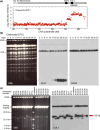
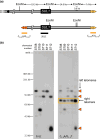
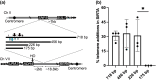
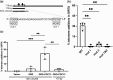
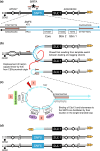
Evolution is driven by the accumulation of competing mutations that influence survival. A broad form of genetic variation is the amplification or deletion of DNA (≥50 bp) referred to as copy number variation (CNV). In humans, CNV may be inconsequential, contribute to minor phenotypic differences, or cause conditions such as birth defects, neurodevelopmental disorders, and cancers. To identify mechanisms that drive CNV, we monitored the experimental evolution of Saccharomyces cerevisiae populations grown under sulfate-limiting conditions. Cells with increased copy number of the gene SUL1, which encodes a primary sulfate transporter, exhibit a fitness advantage. Previously, we reported interstitial inverted triplications of SUL1 as the dominant rearrangement in a haploid population. Here, in a diploid population, we find instead that small linear fragments containing SUL1 form and are sustained over several generations. Many of the linear fragments are stabilized by de novo telomere addition within a telomere-like sequence near SUL1 (within the SNF5 gene). Using an assay that monitors telomerase action following an induced chromosome break, we show that this region acts as a hotspot of de novo telomere addition and that required sequences map to a region of <250 base pairs. Consistent with previous work showing that association of the telomere-binding protein Cdc13 with internal sequences stimulates telomerase recruitment, mutation of a four-nucleotide motif predicted to associate with Cdc13 abolishes de novo telomere addition. Our study suggests that internal telomere-like sequences that stimulate de novo telomere addition can contribute to adaptation by promoting genomic plasticity.
Keywords: Saccharomyces cerevisiae; Cdc13; DNA repair; copy number variation; de novo telomere addition; evolutionary genomics; gene amplification.
© The Author(s) 2023. Published by Oxford University Press on behalf of The Genetics Society of America. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Conflict of interest statement
Conflicts of interest The authors declare no conflict of interest.
Figures

Similar articles
-
A comprehensive map of hotspots of de novo telomere addition in Saccharomyces cerevisiae.Genetics. 2023 May 26;224(2):iyad076. doi: 10.1093/genetics/iyad076. Genetics. 2023. PMID: 37119805 Free PMC article.
-
Interaction of yeast Rad51 and Rad52 relieves Rad52-mediated inhibition of de novo telomere addition.PLoS Genet. 2020 Feb 3;16(2):e1008608. doi: 10.1371/journal.pgen.1008608. eCollection 2020 Feb. PLoS Genet. 2020. PMID: 32012161 Free PMC article.
-
Chromosome healing by de novo telomere addition in Saccharomyces cerevisiae.Mol Microbiol. 2006 Mar;59(5):1357-68. doi: 10.1111/j.1365-2958.2006.05026.x. Mol Microbiol. 2006. PMID: 16468981 Review.
-
Endogenous Hot Spots of De Novo Telomere Addition in the Yeast Genome Contain Proximal Enhancers That Bind Cdc13.Mol Cell Biol. 2016 May 31;36(12):1750-63. doi: 10.1128/MCB.00095-16. Print 2016 Jun 15. Mol Cell Biol. 2016. PMID: 27044869 Free PMC article.
-
Fine tuning the level of the Cdc13 telomere-capping protein for maximal chromosome stability performance.Curr Genet. 2019 Feb;65(1):109-118. doi: 10.1007/s00294-018-0871-3. Epub 2018 Jul 31. Curr Genet. 2019. PMID: 30066139 Review.
Cited by
-
Cdc13 exhibits dynamic DNA strand exchange in the presence of telomeric DNA.Nucleic Acids Res. 2024 Jun 24;52(11):6317-6332. doi: 10.1093/nar/gkae265. Nucleic Acids Res. 2024. PMID: 38613387 Free PMC article.
-
A comprehensive map of hotspots of de novo telomere addition in Saccharomyces cerevisiae.Genetics. 2023 May 26;224(2):iyad076. doi: 10.1093/genetics/iyad076. Genetics. 2023. PMID: 37119805 Free PMC article.
-
A comprehensive map of hotspots of de novo telomere addition in Saccharomyces cerevisiae.bioRxiv [Preprint]. 2023 Mar 23:2023.03.20.533556. doi: 10.1101/2023.03.20.533556. bioRxiv. 2023. Update in: Genetics. 2023 May 26;224(2):iyad076. doi: 10.1093/genetics/iyad076. PMID: 36993206 Free PMC article. Updated. Preprint.
References
-
- Anand R, Memisoglu G, Haber J. Cas9-mediated gene editing in Saccharomyces cerevisiae. Protocol Exchange. 2017. doi:10.1038/protex.2017.021a. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

