QTL mapping and candidate gene analysis for yield and grain weight/size in Tartary buckwheat
- PMID: 36703107
- PMCID: PMC9878770
- DOI: 10.1186/s12870-022-04004-x
QTL mapping and candidate gene analysis for yield and grain weight/size in Tartary buckwheat
Abstract
Background: Grain weight/size influences not only grain yield (GY) but also nutritional and appearance quality and consumer preference in Tartary buckwheat. The identification of quantitative trait loci (QTLs)/genes for grain weight/size is an important objective of Tartary buckwheat genetic research and breeding programs.
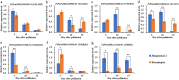
Results: Herein, we mapped the QTLs for GY, 1000-grain weight (TGW), grain length (GL), grain width (GW) and grain length-width ratio (L/W) in four environments using 221 recombinant inbred lines (XJ-RILs) derived from a cross of 'Xiaomiqiao × Jinqiaomai 2'. In total, 32 QTLs, including 7 for GY, 5 for TGW, 6 for GL, 11 for GW and 3 for L/W, were detected and distributed in 24 genomic regions. Two QTL clusters, qClu-1-3 and qClu-1-5, located on chromosome Ft1, were revealed to harbour 7 stable major QTLs for GY (qGY1.2), TGW (qTGW1.2), GL (qGL1.1 and qGL1.4), GW (qGW1.7 and qGW1.10) and L/W (qL/W1.2) repeatedly detected in three and above environments. A total of 59 homologues of 27 known plant grain weight/size genes were found within the physical intervals of qClu-1-3 and qClu-1-5. Six homologues, FtBRI1, FtAGB1, FtTGW6, FtMADS1, FtMKK4 and FtANT, were identified with both non-synonymous SNP/InDel variations and significantly differential expression levels between the two parents, which may play important roles in Tatary buckwheat grain weight/size control and were chosen as core candidate genes for further investigation.
Conclusions: Two stable major QTL clusters related to grain weight/size and six potential key candidate genes were identified by homology comparison, SNP/InDel variations and qRT‒qPCR analysis between the two parents. Our research provides valuable information for improving grain weight/size and yield in Tartary buckwheat breeding.
Keywords: Candidate gene; Grain weight/size; QTL; SNP/InDel variation; Tartary buckwheat; Yield.
© 2023. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
Genetic analysis of yield components in buckwheat using high-throughput sequencing analysis and wild resource populations.Physiol Mol Biol Plants. 2024 Aug;30(8):1313-1328. doi: 10.1007/s12298-024-01491-0. Epub 2024 Jul 22. Physiol Mol Biol Plants. 2024. PMID: 39184561 Free PMC article.
-
QTL Mapping and Candidate Gene Analysis for Starch-Related Traits in Tartary Buckwheat (Fagopyrum tataricum (L.) Gaertn).Int J Mol Sci. 2024 Aug 26;25(17):9243. doi: 10.3390/ijms25179243. Int J Mol Sci. 2024. PMID: 39273191 Free PMC article.
-
Mapping QTLs for 1000-grain weight and genes controlling hull type using SNP marker in Tartary buckwheat (Fagopyrum tataricum).BMC Genomics. 2021 Feb 27;22(1):142. doi: 10.1186/s12864-021-07449-w. BMC Genomics. 2021. PMID: 33639857 Free PMC article.
-
Genetic identification and characterization of quantitative trait loci for wheat grain size-related traits independent of grain number per spike.Theor Appl Genet. 2025 May 25;138(6):125. doi: 10.1007/s00122-025-04912-0. Theor Appl Genet. 2025. PMID: 40413655 Review.
-
Grain Size Associated Genes and the Molecular Regulatory Mechanism in Rice.Int J Mol Sci. 2022 Mar 15;23(6):3169. doi: 10.3390/ijms23063169. Int J Mol Sci. 2022. PMID: 35328589 Free PMC article. Review.
Cited by
-
Genetic analysis of yield components in buckwheat using high-throughput sequencing analysis and wild resource populations.Physiol Mol Biol Plants. 2024 Aug;30(8):1313-1328. doi: 10.1007/s12298-024-01491-0. Epub 2024 Jul 22. Physiol Mol Biol Plants. 2024. PMID: 39184561 Free PMC article.
-
QTL Mapping and Candidate Gene Analysis for Starch-Related Traits in Tartary Buckwheat (Fagopyrum tataricum (L.) Gaertn).Int J Mol Sci. 2024 Aug 26;25(17):9243. doi: 10.3390/ijms25179243. Int J Mol Sci. 2024. PMID: 39273191 Free PMC article.
-
Triumphs of genomic-assisted breeding in crop improvement.Heliyon. 2024 Aug 5;10(15):e35513. doi: 10.1016/j.heliyon.2024.e35513. eCollection 2024 Aug 15. Heliyon. 2024. PMID: 39170454 Free PMC article. Review.
References
MeSH terms
Grants and funding
- QianKeHePingTaiRenCai[2017]5726-18/Training Program of Guizhou Normal University
- (CARS-07-A5)/China Agriculture Research System
- 31860408/Natural Science Foundation of China
- (31960125)/Natural Science Foundation of China
- (2019YFD1001300, 2019YFD1001301)/National Key Research and Development Program of China
LinkOut - more resources
Full Text Sources

