Urine-based multi-omic comparative analysis of COVID-19 and bacterial sepsis-induced ARDS
- PMID: 36703108
- PMCID: PMC9879238
- DOI: 10.1186/s10020-023-00609-6
Urine-based multi-omic comparative analysis of COVID-19 and bacterial sepsis-induced ARDS
Abstract
Background: Acute respiratory distress syndrome (ARDS), a life-threatening condition during critical illness, is a common complication of COVID-19. It can originate from various disease etiologies, including severe infections, major injury, or inhalation of irritants. ARDS poses substantial clinical challenges due to a lack of etiology-specific therapies, multisystem involvement, and heterogeneous, poor patient outcomes. A molecular comparison of ARDS groups holds the potential to reveal common and distinct mechanisms underlying ARDS pathogenesis.
Methods: We performed a comparative analysis of urine-based metabolomics and proteomics profiles from COVID-19 ARDS patients (n = 42) and bacterial sepsis-induced ARDS patients (n = 17). To this end, we used two different approaches, first we compared the molecular omics profiles between ARDS groups, and second, we correlated clinical manifestations within each group with the omics profiles.
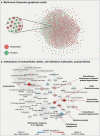
Results: The comparison of the two ARDS etiologies identified 150 metabolites and 70 proteins that were differentially abundant between the two groups. Based on these findings, we interrogated the interplay of cell adhesion/extracellular matrix molecules, inflammation, and mitochondrial dysfunction in ARDS pathogenesis through a multi-omic network approach. Moreover, we identified a proteomic signature associated with mortality in COVID-19 ARDS patients, which contained several proteins that had previously been implicated in clinical manifestations frequently linked with ARDS pathogenesis.
Conclusion: In summary, our results provide evidence for significant molecular differences in ARDS patients from different etiologies and a potential synergy of extracellular matrix molecules, inflammation, and mitochondrial dysfunction in ARDS pathogenesis. The proteomic mortality signature should be further investigated in future studies to develop prediction models for COVID-19 patient outcomes.
Keywords: Acute respiratory distress syndrome (ARDS); COVID-19; Computational analysis; Metabolomics; Mitochondrial dysfunction; Mortality signature; Multi-omic; Network-based; Proteomics.
© 2023. The Author(s).
Conflict of interest statement
A.M.K.C. is a cofounder and equity stockholder for Proterris, which develops therapeutic uses for carbon monoxide. A.M.K.C. has a use patent on CO. Additionally, A.M.K.C. has a patent in COPD. ES consults for Axle informatics regarding COVID vaccine clinical trials through NIAID. JK holds equity in Chymia LLC and IP in PsyProtix and is cofounder of iollo.
Figures


Update of
-
Urine-based multi-omic comparative analysis of COVID-19 and bacterial sepsis-induced ARDS.medRxiv [Preprint]. 2022 Aug 10:2022.08.10.22277939. doi: 10.1101/2022.08.10.22277939. medRxiv. 2022. Update in: Mol Med. 2023 Jan 26;29(1):13. doi: 10.1186/s10020-023-00609-6. PMID: 35982662 Free PMC article. Updated. Preprint.
References
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

