Intrastrand backbone-nucleobase interactions stabilize unwound right-handed helical structures of heteroduplexes of L-aTNA/RNA and SNA/RNA
- PMID: 36703369
- PMCID: PMC9814321
- DOI: 10.1038/s42004-020-00400-2
Intrastrand backbone-nucleobase interactions stabilize unwound right-handed helical structures of heteroduplexes of L-aTNA/RNA and SNA/RNA
Abstract
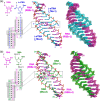
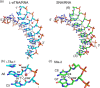
Xeno nucleic acids, which are synthetic analogues of natural nucleic acids, have potential for use in nucleic acid drugs and as orthogonal genetic biopolymers and prebiotic precursors. Although few acyclic nucleic acids can stably bind to RNA and DNA, serinol nucleic acid (SNA) and L-threoninol nucleic acid (L-aTNA) stably bind to them. Here we disclose crystal structures of RNA hybridizing with SNA and with L-aTNA. The heteroduplexes show unwound right-handed helical structures. Unlike canonical A-type duplexes, the base pairs in the heteroduplexes align perpendicularly to the helical axes, and consequently helical pitches are large. The unwound helical structures originate from interactions between nucleobases and neighbouring backbones of L-aTNA and SNA through CH-O bonds. In addition, SNA and L-aTNA form a triplex structure via C:G*G parallel Hoogsteen interactions with RNA. The unique structural features of the RNA-recognizing mode of L-aTNA and SNA should prove useful in nanotechnology, biotechnology, and basic research into prebiotic chemistry.
© 2020. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



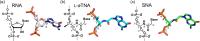

Similar articles
-
Highly stable duplex formation by artificial nucleic acids acyclic threoninol nucleic acid (aTNA) and serinol nucleic acid (SNA) with acyclic scaffolds.Chemistry. 2013 Oct 11;19(42):14151-8. doi: 10.1002/chem.201301578. Epub 2013 Aug 23. Chemistry. 2013. PMID: 24038212
-
Unexpectedly stable homopurine parallel triplex of SNA:RNA*SNA and L-aTNA:RNA*L-aTNA.Chem Commun (Camb). 2024 Jan 30;60(10):1257-1260. doi: 10.1039/d3cc05555h. Chem Commun (Camb). 2024. PMID: 38175608
-
Methyl group configuration on acyclic threoninol nucleic acids (aTNAs) impacts supramolecular properties.Org Biomol Chem. 2022 May 26;20(20):4115-4122. doi: 10.1039/d2ob00266c. Org Biomol Chem. 2022. PMID: 35274662
-
Xeno nucleic acids (XNAs) having non-ribose scaffolds with unique supramolecular properties.Chem Commun (Camb). 2022 Mar 24;58(25):3993-4004. doi: 10.1039/d1cc05868a. Chem Commun (Camb). 2022. PMID: 35107445 Review.
-
Design and Hybridization Properties of Acyclic Xeno Nucleic Acid Oligomers.Chembiochem. 2021 Aug 3;22(15):2507-2515. doi: 10.1002/cbic.202100184. Epub 2021 Jun 2. Chembiochem. 2021. PMID: 33998765 Review.
Cited by
-
Nonenzymatic polymerase-like template-directed synthesis of acyclic L-threoninol nucleic acid.Nat Commun. 2021 Feb 5;12(1):804. doi: 10.1038/s41467-021-21128-0. Nat Commun. 2021. PMID: 33547322 Free PMC article.
-
The XNA alphabet.Nucleic Acids Res. 2025 Jul 8;53(13):gkaf635. doi: 10.1093/nar/gkaf635. Nucleic Acids Res. 2025. PMID: 40650979 Free PMC article. Review.
-
Evaluation of 3'-phosphate as a transient protecting group for controlled enzymatic synthesis of DNA and XNA oligonucleotides.Commun Chem. 2022 Jun 1;5(1):68. doi: 10.1038/s42004-022-00685-5. Commun Chem. 2022. PMID: 36697944 Free PMC article.
-
In vivo efficacy and safety of systemically administered serinol nucleic acid-modified antisense oligonucleotides in mouse kidney.Mol Ther Nucleic Acids. 2024 Dec 18;36(1):102387. doi: 10.1016/j.omtn.2024.102387. eCollection 2025 Mar 11. Mol Ther Nucleic Acids. 2024. PMID: 39850319 Free PMC article.
-
Adenine oligomer directed synthesis of chiral gold nanoparticles.Nat Commun. 2022 Jul 2;13(1):3831. doi: 10.1038/s41467-022-31513-y. Nat Commun. 2022. PMID: 35780141 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

