Gut microbiota signatures in tissues of the colorectal polyp and normal colorectal mucosa, and faeces
- PMID: 36704106
- PMCID: PMC9871776
- DOI: 10.3389/fcimb.2022.1054808
Gut microbiota signatures in tissues of the colorectal polyp and normal colorectal mucosa, and faeces
Abstract
Background: Colorectal polyps are the most common precursors of colorectal cancer (CRC). The close relationship has been observed between colorectal polyps and gut microbiota. However, gut microbiota signatures among sampling sites in patients with colorectal polyps and healthy adults remain elusive.
Aims: To learn about gut microbiota signatures in tissues of the colorectal polyp and normal colorectal mucosa, and faeces.
Methods: We performed 16S rRNA gene sequencing and bioinformatic analysis for the microbiota in the normal colorectal mucosa, the colorectal polyps and faeces of adults with colorectal polyps (n = 24) and in faeces and normal mucosa of healthy adults (n = 16) in this preliminary trial.
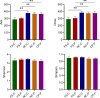
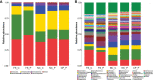
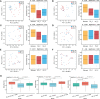
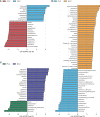
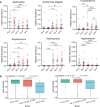
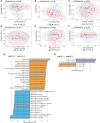
Results: The Ace and Chao indexes were higher in the normal colorectal mucosa and polyp tissues compared to faecal samples (P < 0.05). The composition of microbiota based on PCoA and ANOSIM analysis showed the significant differences only between faeces and tissues of the normal mucosa and polyp (P < 0.05). Based on the LEfSe analysis, the abundances of Bacteroides, Prevotella-2 and Agathobacter were higher, whereas the abundances of Haemophilus, Escherichia_Shigella, Fusobacterium and Streptococcus were lower in faeces both in patients with colorectal polyp and healthy individuals, compared with those in the normal mucosa in two groups or polyp tissues. In healthy individuals, the abundance of Fusobacterium was significantly higher in the normal colorectal mucosa than in faeces. Moreover, there was no significant difference in the abundance of Fusobacterium between the normal colorectal mucosa and polyps in patients with colorectal polyps, but it was significantly higher in the mucosa and polyps than in faeces. Remarkably, the abundance of Fusobacterium in the normal colorectal mucosa was significantly higher in healthy individuals than in the polyp group.
Conclusions: The microbial structure in faeces differs from that in tissues of polyp and normal mucusa. Additionally, Fusobacterium may be a normal colonizer in colonic mucosa, and an abnormal increase of Fusobacterium detected in faeces may be related with the injury of the colorectal mucosa. The difference of the faecal microbiota and mucosal microbiota should be carefully considered in studies on gut microbiota in patients with colorectal lesions.
Keywords: 16S rRNA; Fusobacterium; colorectal polyps; gut microbiota; mucosa.
Copyright © 2023 Zhong, Wang, Xu, Cao, Zhang and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Classification of Changes in the Fecal Microbiota Associated with Colonic Adenomatous Polyps Using a Long-Read Sequencing Platform.Genes (Basel). 2020 Nov 20;11(11):1374. doi: 10.3390/genes11111374. Genes (Basel). 2020. PMID: 33233735 Free PMC article.
-
Microbiome analysis of gut microbiota in patients with colorectal polyps and healthy individuals.Sci Rep. 2025 Feb 28;15(1):7126. doi: 10.1038/s41598-025-91626-4. Sci Rep. 2025. PMID: 40021742 Free PMC article.
-
Study of the Relationship between Microbiome and Colorectal Cancer Susceptibility Using 16SrRNA Sequencing.Biomed Res Int. 2020 Jan 30;2020:7828392. doi: 10.1155/2020/7828392. eCollection 2020. Biomed Res Int. 2020. PMID: 32083132 Free PMC article.
-
Gut microbiota alterations in colorectal adenoma-carcinoma sequence based on 16S rRNA gene sequencing: A systematic review and meta-analysis.Microb Pathog. 2024 Oct;195:106889. doi: 10.1016/j.micpath.2024.106889. Epub 2024 Aug 26. Microb Pathog. 2024. PMID: 39197689
-
Study of Microbiota Associated to Early Tumors Can Shed Light on Colon Carcinogenesis.Int J Mol Sci. 2024 Dec 11;25(24):13308. doi: 10.3390/ijms252413308. Int J Mol Sci. 2024. PMID: 39769073 Free PMC article. Review.
Cited by
-
Gut Microbiota Signatures in Colorectal Cancer as a Potential Diagnostic Biomarker in the Future: A Systematic Review.Int J Mol Sci. 2024 Jul 20;25(14):7937. doi: 10.3390/ijms25147937. Int J Mol Sci. 2024. PMID: 39063179 Free PMC article.
-
Gut Mucosal Microbiome of Patients With Low-Grade Adenomatous Bowel Polyps.Gastro Hep Adv. 2025 Apr 28;4(8):100687. doi: 10.1016/j.gastha.2025.100687. eCollection 2025. Gastro Hep Adv. 2025. PMID: 40530238 Free PMC article.
-
Solobacterium moorei promotes the progression of adenomatous polyps by causing inflammation and disrupting the intestinal barrier.J Transl Med. 2024 Feb 17;22(1):169. doi: 10.1186/s12967-024-04977-3. J Transl Med. 2024. PMID: 38368407 Free PMC article.
-
Dissecting the causal links between gut microbiome, immune traits and polyp using genetic evidence.Front Immunol. 2024 Sep 13;15:1431990. doi: 10.3389/fimmu.2024.1431990. eCollection 2024. Front Immunol. 2024. PMID: 39346904 Free PMC article.
-
Individuality and generality of intratumoral microbiome in the three most prevalent gynecological malignancies: an observational study.Microbiol Spectr. 2024 Sep 3;12(9):e0100424. doi: 10.1128/spectrum.01004-24. Epub 2024 Aug 5. Microbiol Spectr. 2024. PMID: 39101825 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

