Role of the blue light receptor gene Icwc-1 in mycelium growth and fruiting body formation of Isaria cicadae
- PMID: 36704565
- PMCID: PMC9871644
- DOI: 10.3389/fmicb.2022.1038034
Role of the blue light receptor gene Icwc-1 in mycelium growth and fruiting body formation of Isaria cicadae
Abstract
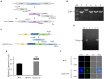
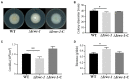
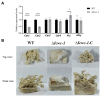
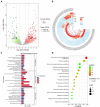
The Isaria cicadae, is well known highly prized medicinal mushroom with great demand in food and pharmaceutical industry. Due to its economic value and therapeutic uses, natural sources of wild I. cicadae are over-exploited and reducing continuously. Therefore, commercial cultivation in controlled environment is an utmost requirement to fulfill the consumer's demand. Due to the lack of knowledge on fruiting body (synnemata) development and regulation, commercial cultivation is currently in a difficult situation. In the growth cycle of macrofungi, such as mushrooms, light is the main factor affecting growth and development, but so far, specific effects of light on the growth and development of I. cicadae is unknown. In this study, we identified a blue light receptor white-collar-1 (Icwc-1) gene homologue with well-defined functions in morphological development in I. cicadae based on gene knockout technology and transcriptomic analysis. It was found that the Icwc-1 gene significantly affected hyphal growth and fruiting body development. This study confirms that Icwc-1 acts as an upstream regulatory gene that regulates genes associated with fruiting body formation, pigment-forming genes, and related genes for enzyme synthesis. Transcriptome data analysis also found that Icwc-1 affects many important metabolic pathways of I. cicadae, i.e., amino acid metabolism and fatty acid metabolism. The above findings will not only provide a comprehensive understanding about the molecular mechanism of light regulation in I. cicadae, but also provide new insights for future breeding program and improving this functional food production.
Keywords: Isaria cicadae; blue light receptor; developmental regulation; fruiting body formation; mycelium growth.
Copyright © 2023 Song, Shrivastava, Gai, Li, Cai, Shen, Lin, Liu and Wang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Effect of Environmental Conditions on Synnema Formation and Nucleoside Production in Cicada Flower, Isaria cicadae (Ascomycetes).Int J Med Mushrooms. 2019;21(1):59-69. doi: 10.1615/IntJMedMushrooms.2018029506. Int J Med Mushrooms. 2019. PMID: 30806256
-
GPI-Anchored Protein Homolog IcFBR1 Functions Directly in Morphological Development of Isaria cicadae.J Fungi (Basel). 2022 Oct 31;8(11):1152. doi: 10.3390/jof8111152. J Fungi (Basel). 2022. PMID: 36354919 Free PMC article.
-
Omics data reveal the unusual asexual-fruiting nature and secondary metabolic potentials of the medicinal fungus Cordyceps cicadae.BMC Genomics. 2017 Aug 30;18(1):668. doi: 10.1186/s12864-017-4060-4. BMC Genomics. 2017. PMID: 28854898 Free PMC article.
-
Can Cordyceps cicadae be used as an alternative to Cordyceps militaris and Cordyceps sinensis? - A review.J Ethnopharmacol. 2020 Jul 15;257:112879. doi: 10.1016/j.jep.2020.112879. Epub 2020 Apr 16. J Ethnopharmacol. 2020. PMID: 32305637 Review.
-
Secondary metabolites (SMs) of Isaria cicadae and Isaria tenuipes.RSC Adv. 2018 Dec 21;9(1):172-184. doi: 10.1039/c8ra09039d. eCollection 2018 Dec 19. RSC Adv. 2018. PMID: 35521576 Free PMC article. Review.
Cited by
-
Roles of CcDFR and CcOMT9 in the cyanidin biosynthesis and development of Cordyceps cicadae.Front Microbiol. 2024 Mar 6;15:1353710. doi: 10.3389/fmicb.2024.1353710. eCollection 2024. Front Microbiol. 2024. PMID: 38511011 Free PMC article.
-
Current Advances in the Functional Genes of Edible and Medicinal Fungi: Research Techniques, Functional Analysis, and Prospects.J Fungi (Basel). 2024 Apr 25;10(5):311. doi: 10.3390/jof10050311. J Fungi (Basel). 2024. PMID: 38786666 Free PMC article. Review.
-
Research hotspots and emerging trends in growth and development of macrofungi: a bibliometric review based on CiteSpace analysis.World J Microbiol Biotechnol. 2024 Oct 26;40(11):365. doi: 10.1007/s11274-024-04168-8. World J Microbiol Biotechnol. 2024. PMID: 39455463 Review.
References
LinkOut - more resources
Full Text Sources

