Direct lysis of 3D cell cultures for RT-qPCR gene expression quantification
- PMID: 36707637
- PMCID: PMC9883454
- DOI: 10.1038/s41598-023-28844-1
Direct lysis of 3D cell cultures for RT-qPCR gene expression quantification
Abstract
In vitro cell culture experiments are widely used to study cellular behavior in most biological research fields. Except for suspension cells, most human cell types are cultured as adherent monolayers on a plastic surface. While technically convenient, monolayer cultures can suffer from limitations in terms of physiological relevance, as their resemblance to complex in vivo tissue structures is limited. To address these limitations, three-dimensional (3D) cell culture systems have gained increased interest as they mimic key structural and functional properties of their in vivo tissue counterparts. Nevertheless, protocols established on monolayer cell cultures may require adjustments if they are to be applied to 3D cell cultures. As gene expression quantification is an essential part of many in vitro experiments, we evaluated and optimized a direct cell lysis, reverse transcription and qPCR protocol applicable for 3D cell cultures. The newly developed protocol wherein gene expression is determined directly from crude cell lysates showed improved cell lysis compared to the standard protocol, accurate gene expression quantification, hereby avoiding time-consuming cell harvesting and RNA extraction.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
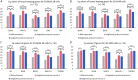
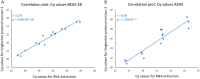
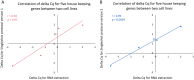
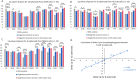
Similar articles
-
Implications of adipose-derived stromal cells in a 3D culture system for osteogenic differentiation: an in vitro and in vivo investigation.Spine J. 2013 Jan;13(1):32-43. doi: 10.1016/j.spinee.2013.01.002. Spine J. 2013. PMID: 23384881
-
Validation of a PicoGreen-based DNA quantification integrated in an RNA extraction method for two-dimensional and three-dimensional cell cultures.Tissue Eng Part C Methods. 2012 Jun;18(6):444-52. doi: 10.1089/ten.TEC.2011.0304. Epub 2012 Jan 26. Tissue Eng Part C Methods. 2012. PMID: 22195986
-
Dynamic three-dimensional culture methods enhance mesenchymal stem cell properties and increase therapeutic potential.Tissue Eng Part C Methods. 2010 Aug;16(4):735-49. doi: 10.1089/ten.TEC.2009.0432. Tissue Eng Part C Methods. 2010. PMID: 19811095
-
Cancer Cytokines and the Relevance of 3D Cultures for Studying Those Implicated in Human Cancers.J Cell Biochem. 2017 Sep;118(9):2544-2558. doi: 10.1002/jcb.25970. Epub 2017 May 3. J Cell Biochem. 2017. PMID: 28262975 Review.
-
Devices and techniques used to obtain and analyze three-dimensional cell cultures.Biotechnol Prog. 2021 May;37(3):e3126. doi: 10.1002/btpr.3126. Epub 2021 Jan 29. Biotechnol Prog. 2021. PMID: 33460298 Review.
Cited by
-
Sequential orthogonal assays for longitudinal and endpoint characterization of three-dimensional spheroids.Nat Protoc. 2025 Apr 8. doi: 10.1038/s41596-025-01150-y. Online ahead of print. Nat Protoc. 2025. PMID: 40200041 Review.
References
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

