Placental circadian lincRNAs and spontaneous preterm birth
- PMID: 36712876
- PMCID: PMC9874002
- DOI: 10.3389/fgene.2022.1051396
Placental circadian lincRNAs and spontaneous preterm birth
Abstract
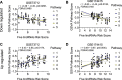
Long non-coding RNAs (lncRNAs) have a much higher cell- and/or tissue-specificity compared to mRNAs in most cases, making them excellent candidates for therapeutic applications to reduce off-target effects. Placental long non-coding RNAs have been investigated in the pathogenesis of preeclampsia (often causing preterm birth (PTB)), but less is known about their role in preterm birth. Preterm birth occurs in 11% of pregnancies and is the most common cause of death among infants in the world. We recently identified that genes that drive circadian rhythms in cells, termed molecular clock genes, are deregulated in maternal blood of women with spontaneous PTB (sPTB) and in the placenta of women with preeclampsia. Next, we focused on circadian genes-correlated long intergenic non-coding RNAs (lincRNAs, making up most of the long non-coding RNAs), designated as circadian lincRNAs, associated with sPTB. We compared the co-altered circadian transcripts-correlated lincRNAs expressed in placentas of sPTB and term births using two published independent RNAseq datasets (GSE73712 and GSE174415). Nine core clock genes were up- or downregulated in sPTB versus term birth, where the RORA transcript was the only gene downregulated in sPTB across both independent datasets. We found that five circadian lincRNAs (LINC00893, LINC00265, LINC01089, LINC00482, and LINC00649) were decreased in sPTB vs term births across both datasets (p ≤ .0222, FDR≤.1973) and were negatively correlated with the dataset-specific clock genes-based risk scores (correlation coefficient r = -.65 ∼ -.43, p ≤ .0365, FDR≤.0601). Gene set variation analysis revealed that 65 pathways were significantly enriched by these same five differentially expressed lincRNAs, of which over 85% of the pathways could be linked to immune/inflammation/oxidative stress and cell cycle/apoptosis/autophagy/cellular senescence. These findings may improve our understanding of the pathogenesis of spontaneous preterm birth and provide novel insights into the development of potentially more effective and specific therapeutic targets against sPTB.
Keywords: GSVA analysis; lincRNA; molecular clock; preterm (birth); transcriptional co-alteration.
Copyright © 2023 Zhou, Fichorova, Holzman, Chen, Chang, Kasten and Hoffmann.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


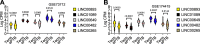


References
-
- Ackerman W. E., Buhimschi I. A., Eidem H. R., Rinker D. C., Rokas A., Rood K., et al. (2016). Comprehensive RNA profiling of villous trophoblast and decidua basalis in pregnancies complicated by preterm birth following intra-amniotic infection. Placenta 44, 23–33. 10.1016/j.placenta.2016.05.010 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

