A clinician's guide to understanding and critically appraising machine learning studies: a checklist for Ruling Out Bias Using Standard Tools in Machine Learning (ROBUST-ML)
- PMID: 36713011
- PMCID: PMC9708024
- DOI: 10.1093/ehjdh/ztac016
A clinician's guide to understanding and critically appraising machine learning studies: a checklist for Ruling Out Bias Using Standard Tools in Machine Learning (ROBUST-ML)
Abstract
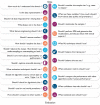
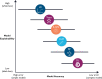
Developing functional machine learning (ML)-based models to address unmet clinical needs requires unique considerations for optimal clinical utility. Recent debates about the rigours, transparency, explainability, and reproducibility of ML models, terms which are defined in this article, have raised concerns about their clinical utility and suitability for integration in current evidence-based practice paradigms. This featured article focuses on increasing the literacy of ML among clinicians by providing them with the knowledge and tools needed to understand and critically appraise clinical studies focused on ML. A checklist is provided for evaluating the rigour and reproducibility of the four ML building blocks: data curation, feature engineering, model development, and clinical deployment. Checklists like this are important for quality assurance and to ensure that ML studies are rigourously and confidently reviewed by clinicians and are guided by domain knowledge of the setting in which the findings will be applied. Bridging the gap between clinicians, healthcare scientists, and ML engineers can address many shortcomings and pitfalls of ML-based solutions and their potential deployment at the bedside.
Keywords: Bias; Critical appraisal; Guidelines; Healthcare; Machine learning; Quality.
© The Author(s) 2022. Published by Oxford University Press on behalf of the European Society of Cardiology.
Figures






References
-
- Leisman DE, Harhay MO, Lederer DJ, Abramson M, Adjei AA, Bakker J, Ballas ZK, Barreiro E, Bell SC, Bellomo R, Bernstein JA, Branson RD, Brusasco V, Chalmers JD, Chokroverty S, Citerio G, Collop NA, Cooke CR, Crapo JD, Donaldson G, Fitzgerald DA, Grainger E, Hale L, Herth FJ, Kochanek PM, Marks G, Moorman JR, Ost DE, Schatz M, Sheikh A, Smyth AR, Stewart I, Stewart PW, Swenson ER, Szymusiak R, Teboul J-L, Vincent J-L, Wedzicha JA, Maslove DM. Development and reporting of prediction models: guidance for authors from editors of respiratory, sleep, and critical care journals. Crit Care Med 2020;48:623–633. - PMC - PubMed
-
- Rajkomar A, Dean J, Kohane I. Machine Learning in Medicine. N Engl J Med 2019;380:1347–1358. - PubMed
-
- Al’Aref SJ, Anchouche K, Singh G, Slomka PJ, Kolli KK, Kumar A, Pandey M, Maliakal G, Van Rosendael AR, Beecy AN, Berman DS, Leipsic J, Nieman K, Andreini D, Pontone G, Schoepf UJ, Shaw LJ, Chang H-J, Narula J, Bax JJ, Guan Y, Min JK. Clinical applications of machine learning in cardiovascular disease and its relevance to cardiac imaging. Eur Heart J 2019;40:1975–1986. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous
