Annotation of G-Protein Coupled Receptors in the Genomes of Parasitic Blood Flukes
- PMID: 36713056
- PMCID: PMC9874797
- DOI: 10.17912/micropub.biology.000704
Annotation of G-Protein Coupled Receptors in the Genomes of Parasitic Blood Flukes
Abstract
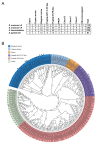
Infection with Schistosoma parasitic flatworms ( Schistosoma haematobium, Schistosoma mansoni and Schistosoma japonicum ) causes the neglected tropical disease schistosomiasis. There is a need to identify new chemotherapies to treat these parasites, and G-protein coupled receptors (GPCRs) are a logical druggable targets to explore given they control key aspects of schistosome biology such as neuromuscular function and reproduction. Updated chromosome level genome assemblies for each of the three major species have recently been released. However, studies on these GPCRs require accurate, updated genome annotations. Here, we have re-annotated the GPCRs present in each of the three major schistosome species.
Copyright: © 2023 by the authors.
Figures
References
-
- Berriman M, Haas BJ, LoVerde PT, Wilson RA, Dillon GP, Cerqueira GC, Mashiyama ST, Al-Lazikani B, Andrade LF, Ashton PD, Aslett MA, Bartholomeu DC, Blandin G, Caffrey CR, Coghlan A, Coulson R, Day TA, Delcher A, DeMarco R, Djikeng A, Eyre T, Gamble JA, Ghedin E, Gu Y, Hertz-Fowler C, Hirai H, Hirai Y, Houston R, Ivens A, Johnston DA, Lacerda D, Macedo CD, McVeigh P, Ning Z, Oliveira G, Overington JP, Parkhill J, Pertea M, Pierce RJ, Protasio AV, Quail MA, Rajandream MA, Rogers J, Sajid M, Salzberg SL, Stanke M, Tivey AR, White O, Williams DL, Wortman J, Wu W, Zamanian M, Zerlotini A, Fraser-Liggett CM, Barrell BG, El-Sayed NM. The genome of the blood fluke Schistosoma mansoni. Nature. 2009 Jul 16;460(7253):352–358. doi: 10.1038/nature08160. - DOI - PMC - PubMed
-
- Buddenborg Sarah K, Tracey Alan, Berger Duncan J, Lu Zhigang, Doyle Stephen R, Fu Beiyuan, Yang Fengtang, Reid Adam J, Rodgers Faye H, Rinaldi Gabriel, Sankaranarayanan Geetha, Böhme Ulrike, Holroyd Nancy, Berriman Matthew. 2021 Aug 13; doi: 10.1101/2021.08.13.456314. - DOI
LinkOut - more resources
Full Text Sources
