Fourth Report on Chicken Genes and Chromosomes 2022
- PMID: 36716736
- PMCID: PMC11835228
- DOI: 10.1159/000529376
Fourth Report on Chicken Genes and Chromosomes 2022
Conflict of interest statement
All authors have no conflicts of interest.
The authors have no conflicts of interest to declare.
There are no conflicts of interest to declare.
The authors declare no conflicts of interest.
The authors declare no competing interests.
The authors have no conflicts of interest to declare.
There are no conflicts of interest.
The authors have no conflicts of interest to declare.
The authors have no conflicts of interest to declare.
The authors report no competing interests.
The authors declare that they have no competing interests.
The author has no conflicts of interest to declare.
The authors have no conflicts of interest to declare.
The author declares no conflict of interest.
The author declares no conflicts of interest.
The authors declare no conflicts of interest.
The authors declare that they have no conflict of interest.
The authors have no conflicts of interest to declare.
The authors of this document declare that they have no affiliations with or involvement in any organization or entity with any financial interest in the subject matter or materials discussed in this manuscript.
The authors have no conflicts of interest to declare.
The authors have no conflicts of interest to declare.
Figures

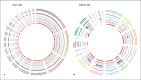

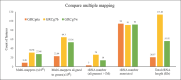

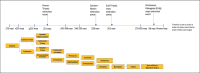


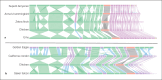
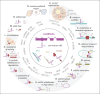

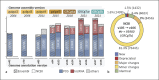
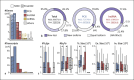
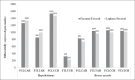
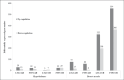
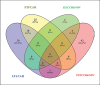
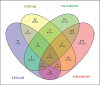

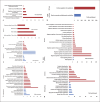



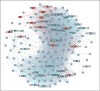
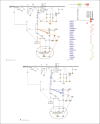


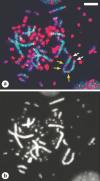


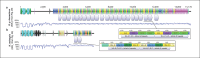


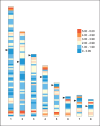
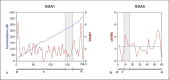
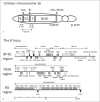
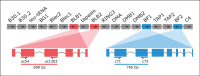

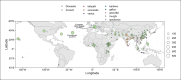
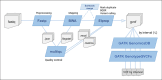



References
-
- Aarts HJ, Van Der Hulst-Van Arkel MC, Beuving G, Leenstra FR. Variations in endogenous viral gene patterns in white leghorn, medium heavy, white Plymouth Rock, and Cornish chickens. Poult Sci. 1991;70((6)):1281–6. - PubMed
-
- Abdelnour SA, Abd El-Hack ME, Khafaga AF, Arif M, Taha AE, Noreldin AE. Stress biomarkers and proteomics alteration to thermal stress in ruminants: a review. J Therm Biol. 2019;79:120–34. - PubMed
Publication types
LinkOut - more resources
Full Text Sources

