Food protein-derived amyloids do not accelerate amyloid β aggregation
- PMID: 36720893
- PMCID: PMC9889329
- DOI: 10.1038/s41598-023-28147-5
Food protein-derived amyloids do not accelerate amyloid β aggregation
Abstract
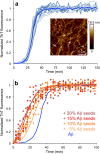
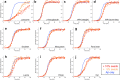
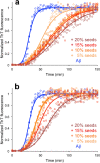
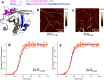
The deposition of proteins in the form of amyloid fibrils is closely associated with several serious diseases. The events that trigger the conversion from soluble functional proteins into insoluble amyloid are not fully understood. Many proteins that are not associated with disease can form amyloid with similar structural characteristics as the disease-associated fibrils, which highlights the potential risk of cross-seeding of disease amyloid by amyloid-like structures encountered in our surrounding. Of particular interest are common food proteins that can be transformed into amyloid under conditions similar to cooking. We here investigate cross-seeding of amyloid-β (Aβ), a peptide known to form amyloid during the development of Alzheimer's disease, by 16 types of amyloid fibrils derived from food proteins or peptides. Kinetic studies using thioflavin T fluorescence as output show that none of the investigated protein fibrils accelerates the aggregation of Aβ. In at least two cases (hen egg lysozyme and oat protein isolate) we observe retardation of the aggregation, which appears to originate from interactions between the food protein seeds and Aβ in aggregated form. The results support the view that food-derived amyloid is not a risk factor for development of Aβ pathology and Alzheimer's disease.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

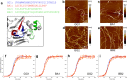
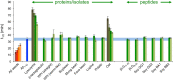
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

