Systematic comparison of HIV-1 Envelope-specific IgG responses induced by different vaccination regimens: Can we steer IgG recognition towards regions of viral vulnerability?
- PMID: 36741409
- PMCID: PMC9891136
- DOI: 10.3389/fimmu.2022.1075606
Systematic comparison of HIV-1 Envelope-specific IgG responses induced by different vaccination regimens: Can we steer IgG recognition towards regions of viral vulnerability?
Abstract
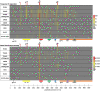
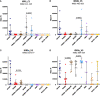
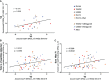
Immunogens and vaccination regimens can influence patterns of immune-epitope recognition, steering them towards or away from epitopes of potential viral vulnerability. HIV-1 envelope (Env)-specific antibodies targeting variable region 2 (V2) or 3 (V3) correlated with protection during the RV144 trial, however, it was suggested that the immunodominant V3 region might divert antibody responses away from other relevant sites. We mapped IgG responses against linear Env epitopes in five clinical HIV vaccine trials, revealing a specific pattern of Env targeting for each regimen. Notable V2 responses were only induced in trials administering CRF01_AE based immunogens, but targeting of V3 was seen in all trials, with the soluble, trimeric CN54gp140 protein eliciting robust V3 recognition. Strong V3 targeting was linked to greater overall response, increased number of total recognised antigenic regions, and where present, stronger V2 recognition. Hence, strong induction of V3-specific antibodies did not negatively impact the targeting of other linear epitopes in this study, suggesting that the induction of antibodies against V3 and other regions of potential viral vulnerability need not be necessarily mutually exclusive.
Keywords: CN54rgp140 vaccine; HIV; V2-antibodies; V3-antibodies; envelope-specific antibodies; immunogen sequence; linear peptide array; vaccine.
Copyright © 2023 Horvath, Rogers, Pollakis, Baranov, Pieroth, Joseph, Chachage, Heitzer, Maganga, Msafiri, Joachim, Viegas, Eller, Kibuuka, Rerks-Ngarm, Pitisuttithum, Nitayapan, Dhitavat, Premsri, Fidler, Shattock, Robb, Weber, McCormack, Munseri, Lyamuya, Nilsson, Kroidl, Hoelscher, Wagner, Geldmacher and Held.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

