Identification and characterization of plasmids carrying the mobile colistin resistance gene mcr-1 using optical DNA mapping
- PMID: 36743530
- PMCID: PMC9891347
- DOI: 10.1093/jacamr/dlad004
Identification and characterization of plasmids carrying the mobile colistin resistance gene mcr-1 using optical DNA mapping
Abstract
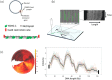
Objectives: Colistin is a last-resort antibiotic, but there has been a rapid increase in colistin resistance, threatening its use in the treatment of infections with carbapenem-resistant Enterobacterales (CRE). Plasmid-mediated colistin resistance, in particular the mcr-1 gene, has been identified and WGS is the go-to method in identifying plasmids carrying mcr-1 genes. The goal of this study is to demonstrate the use of optical DNA mapping (ODM), a fast, efficient and amplification-free technique, to characterize plasmids carrying mcr-1.
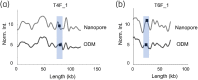
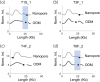
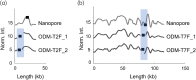
Methods: ODM is a single-molecule technique, which we have demonstrated can be used for identifying plasmids harbouring antibiotic resistance genes. We here applied the technique to plasmids isolated from 12 clinical Enterobacterales isolates from patients at a major hospital in Thailand and verified our results using Nanopore long-read sequencing.
Results: We successfully identified plasmids encoding the mcr-1 gene and, for the first time, demonstrated the ability of ODM to identify resistance gene sites in small (∼30 kb) plasmids. We further identified bla CTX-M genes in different plasmids than the ones encoding mcr-1 in three of the isolates studied. Finally, we propose a cut-and-stretch assay, based on similar principles, but performed using surface-functionalized cover slips for DNA immobilization and an inexpensive microscope with basic functionalities, to identify the mcr-1 gene in a plasmid sample.
Conclusions: Both ODM and the cut-and-stretch assay developed could be very useful in identifying plasmids encoding antibiotic resistance in hospitals and healthcare facilities. The cut-and-stretch assay is particularly useful in low- and middle-income countries, where existing techniques are limited.
© The Author(s) 2023. Published by Oxford University Press on behalf of British Society for Antimicrobial Chemotherapy.
Figures
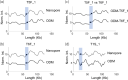

References
LinkOut - more resources
Full Text Sources
