Transcriptomic and metabolic regulatory network characterization of drought responses in tobacco
- PMID: 36743571
- PMCID: PMC9891310
- DOI: 10.3389/fpls.2022.1067076
Transcriptomic and metabolic regulatory network characterization of drought responses in tobacco
Abstract
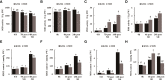
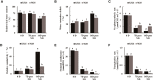
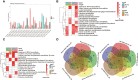
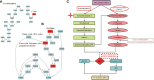
Drought stress usually causes huge economic losses for tobacco industries. Drought stress exhibits multifaceted impacts on tobacco systems through inducing changes at different levels, such as physiological and chemical changes, changes of gene transcription and metabolic changes. Understanding how plants respond and adapt to drought stress helps generate engineered plants with enhanced drought resistance. In this study, we conducted multiple time point-related physiological, biochemical,transcriptomic and metabolic assays using K326 and its derived mutant 28 (M28) with contrasting drought tolerance. Through integrative analyses of transcriptome and metabolome,we observed dramatic changes of gene expression and metabolic profiles between M28 and K326 before and after drought treatment. we found that some of DEGs function as key enzymes responsible for ABA biosynthesis and metabolic pathway, thereby mitigating impairment of drought stress through ABA signaling dependent pathways. Four DEGs were involved in nitrogen metabolism, leading to synthesis of glutamate (Glu) starting from NO-3 /NO-2 that serves as an indicator for stress responses. Importantly, through regulatory network analyses, we detected several drought induced TFs that regulate expression of genes responsible for ABA biosynthesis through network, indicating direct and indirect involvement of TFs in drought responses in tobacco. Thus, our study sheds some mechanistic insights into how plant responding to drought stress through transcriptomic and metabolic changes in tobacco. It also provides some key TF or non-TF gene candidates for engineering manipulation for breeding new tobacco varieties with enhanced drought tolerance.
Keywords: Nicotiana tabacum L.; drought stress; metabolome; regulatory network; transcriptome.
Copyright © 2023 Hu, He, Li, Wang, Yi, Yang, Yang, Borovskii, Cheng, Hu and Zhang.
Conflict of interest statement
Author PY was employed by company Hu'nan Tobacco Company Changde Company, Changde, Hunan, China. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



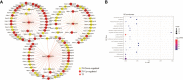
References
LinkOut - more resources
Full Text Sources
Miscellaneous

