The organizer of chromatin topology RIF1 ensures cellular resilience to DNA replication stress
- PMID: 36746532
- PMCID: PMC9906048
- DOI: 10.26508/lsa.202101186
The organizer of chromatin topology RIF1 ensures cellular resilience to DNA replication stress
Abstract
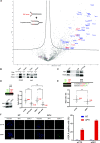
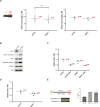
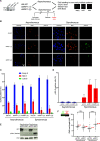
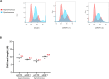
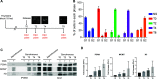
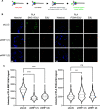
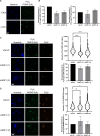
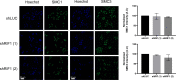
Eukaryotic genomes are duplicated from thousands of replication origins that fire sequentially forming a defined spatiotemporal pattern of replication clusters. The temporal order of DNA replication is determined by chromatin architecture and, more specifically, by chromatin contacts that are stabilized by RIF1. Here, we show that RIF1 localizes near newly synthesized DNA. In cells exposed to the DNA replication inhibitor aphidicolin, suppression of RIF1 markedly decreased the efficacy of isolation of proteins on nascent DNA, suggesting that the isolation of proteins on nascent DNA procedure is biased by chromatin topology. RIF1 was required to limit the accumulation of DNA lesions induced by aphidicolin treatment and promoted the recruitment of cohesins in the vicinity of nascent DNA. Collectively, the data suggest that the stabilization of chromatin topology by RIF1 limits replication-associated genomic instability.
© 2023 Lebdy et al.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures




References
-
- Alabert C, Bukowski-Wills JC, Lee SB, Kustatscher G, Nakamura K, de Lima Alves F, Menard P, Mejlvang J, Rappsilber J, Groth A (2014) Nascent chromatin capture proteomics determines chromatin dynamics during DNA replication and identifies unknown fork components. Nat Cell Biol 16: 281–291. 10.1038/ncb2918 - DOI - PMC - PubMed
-
- Balasubramanian S, Andreani M, Andrade JG, Saha T, Sundaravinayagam D, Garzon J, Zhang W, Popp O, Hiraga SI, Rahjouei A, et al. (2022) Protection of nascent DNA at stalled replication forks is mediated by phosphorylation of RIF1 intrinsically disordered region. Elife 11: e75047. 10.7554/elife.75047 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous
