Programmable synthetic biomolecular condensates for cellular control
- PMID: 36747054
- PMCID: PMC10786170
- DOI: 10.1038/s41589-022-01252-8
Programmable synthetic biomolecular condensates for cellular control
Abstract
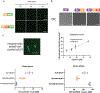
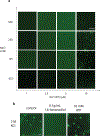
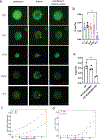
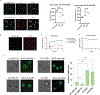
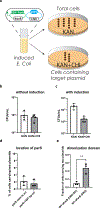
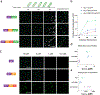
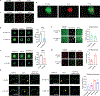
The formation of biomolecular condensates mediated by a coupling of associative and segregative phase transitions plays a critical role in controlling diverse cellular functions in nature. This has inspired the use of phase transitions to design synthetic systems. While design rules of phase transitions have been established for many synthetic intrinsically disordered proteins, most efforts have focused on investigating their phase behaviors in a test tube. Here, we present a rational engineering approach to program the formation and physical properties of synthetic condensates to achieve intended cellular functions. We demonstrate this approach through targeted plasmid sequestration and transcription regulation in bacteria and modulation of a protein circuit in mammalian cells. Our approach lays the foundation for engineering designer condensates for synthetic biology applications.
© 2023. The Author(s), under exclusive licence to Springer Nature America, Inc.
Conflict of interest statement
COMPETING INTERESTS STATEMENT
The authors declare no competing interesting for this work.
Figures
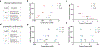




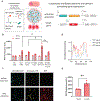

References
-
- Brangwynne CP et al. Germline P Granules Are Liquid Droplets That Localize by Controlled Dissolution/Condensation. Science 324, 1729–1732 (2009). - PubMed
-
- Tsang B, Pritišanac I, Scherer SW, Moses AM & Forman-Kay JD Phase Separation as a Missing Mechanism for Interpretation of Disease Mutations. Cell 183, 1742–1756 (2020). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

