Preclinical evaluation of a COVID-19 vaccine candidate based on a recombinant RBD fusion heterodimer of SARS-CoV-2
- PMID: 36748086
- PMCID: PMC9893798
- DOI: 10.1016/j.isci.2023.106126
Preclinical evaluation of a COVID-19 vaccine candidate based on a recombinant RBD fusion heterodimer of SARS-CoV-2
Abstract
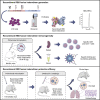
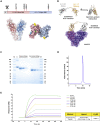
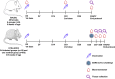
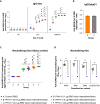
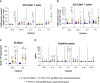
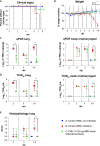
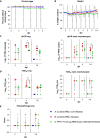
Current COVID-19 vaccines have been associated with a decline in infection rates, prevention of severe disease, and a decrease in mortality rates. However, SARS-CoV-2 variants are continuously evolving, and development of new accessible COVID-19 vaccines is essential to mitigate the pandemic. Here, we present data on preclinical studies in mice of a receptor-binding domain (RBD)-based recombinant protein vaccine (PHH-1V) consisting of an RBD fusion heterodimer comprising the B.1.351 and B.1.1.7 SARS-CoV-2 variants formulated in SQBA adjuvant, an oil-in-water emulsion. A prime-boost immunisation with PHH-1V in BALB/c and K18-hACE2 mice induced a CD4+ and CD8+ T cell response and RBD-binding antibodies with neutralizing activity against several variants, and also showed a good tolerability profile. Significantly, RBD fusion heterodimer vaccination conferred 100% efficacy, preventing mortality in SARS-CoV-2 infected K18-hACE2 mice, but also reducing Beta, Delta and Omicron infection in lower respiratory airways. These findings demonstrate the feasibility of this recombinant vaccine strategy.
Keywords: Health sciences; Immune response; Immunology; Microbiology.
© 2023 The Author(s).
Conflict of interest statement
Authors indicated as “1” are employees of HIPRA, a private pharmaceutical company that develops and manufactures vaccines. CReSA, IrsiCaixa, CMCiB-IGTP, UPF, and ICREA have received financial support from HIPRA. Two patent applications have been filed by HIPRA Scientific S.L.U. and Laboratorios HIPRA, S.A., on different SARS-CoV-2 vaccine candidates and SARS-CoV-2 subunit vaccines, including the recombinant RBD fusion heterodimer PHH-1V. Antonio Barreiro, Antoni Prenafeta, Luis González, Laura Ferrer, Ester Puigvert, Jordi Palmada, Teresa Prat, and Carme Garriga are the inventors of these patent applications.
Figures


References
-
- WHO . Word Health Organization; 2021. WHO Coronavirus (COVID-19) Dashboard. 2021.https://covid19.who.int/
-
- Ritchie H., Rodés-Guirao L., Appel C., Giattino C., Ortiz-Ospina E., Hasell J., Macdonald B., Beltekian D., Roser M. Coronavirus Pandemic (COVID-19) 2022. https://ourworldindata.org/coronavirus
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

