Learning Correlations between Internal Coordinates to Improve 3D Cartesian Coordinates for Proteins
- PMID: 36749957
- PMCID: PMC10404647
- DOI: 10.1021/acs.jctc.2c01270
Learning Correlations between Internal Coordinates to Improve 3D Cartesian Coordinates for Proteins
Abstract
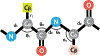
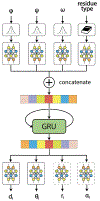
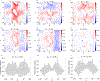
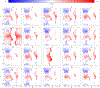
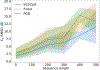
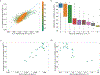
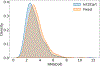
We consider a generic representation problem of internal coordinates (bond lengths, valence angles, and dihedral angles) and their transformation to 3-dimensional Cartesian coordinates of a biomolecule. We show that the internal-to-Cartesian process relies on correctly predicting chemically subtle correlations among the internal coordinates themselves, and learning these correlations increases the fidelity of the Cartesian representation. We developed a machine learning algorithm, Int2Cart, to predict bond lengths and bond angles from backbone torsion angles and residue types of a protein, which allows reconstruction of protein structures better than using fixed bond lengths and bond angles or a static library method that relies on backbone torsion angles and residue types in a local environment. The method is able to be used for structure validation, as we show that the agreement between Int2Cart-predicted bond geometries and those from an AlphaFold 2 model can be used to estimate model quality. Additionally, by using Int2Cart to reconstruct an IDP ensemble, we are able to decrease the clash rate during modeling. The Int2Cart algorithm has been implemented as a publicly accessible python package at https://github.com/THGLab/int2cart.
Figures

References
-
- Baker J; Kinghorn D; Pulay P. Geometry optimization in delocalized internal coordinates: An efficient quadratically scaling algorithm for large molecules. J. Chem. Phys 1999, 110, 4986–4991.
-
- Schwieters CD; Clore G. Internal Coordinates for Molecular Dynamics and Minimization in Structure Determination and Refinement. J. Magn. Reson 2001, 152, 288–302. - PubMed
-
- Parsons J; Holmes JB; Rojas JM; Tsai J; Strauss CE Practical conversion from torsion space to Cartesian space for in silico protein synthesis. J. Comput. Chem 2005, 26, 1063–1068. - PubMed
