SCARECROW-like GRAS protein PES positively regulates petunia floral scent production
- PMID: 36760164
- PMCID: PMC10152688
- DOI: 10.1093/plphys/kiad081
SCARECROW-like GRAS protein PES positively regulates petunia floral scent production
Abstract
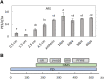
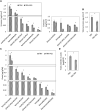
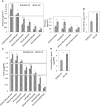
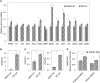
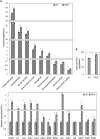
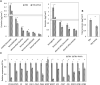
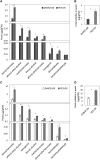
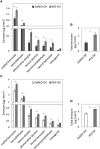
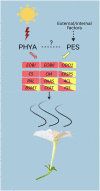
Emission of scent volatiles by flowers is important for successful pollination and consequently, reproduction. Petunia (Petunia hybrida) floral scent is formed mainly by volatile products of the phenylpropanoid pathway. We identified and characterized a regulator of petunia scent production: the GRAS protein PHENYLPROPANOID EMISSION-REGULATING SCARECROW-LIKE (PES). Its expression increased in petals during bud development and was highest in open flowers. Overexpression of PES increased the production of floral volatiles, while its suppression resulted in scent reduction. We showed that PES upregulates the expression of genes encoding enzymes of the phenylpropanoid and shikimate pathways in petals, and of the core regulator of volatile biosynthesis ODORANT1 by activating its promoter. PES is an ortholog of Arabidopsis (Arabidopsis thaliana) PHYTOCHROME A SIGNAL TRANSDUCTION 1, involved in physiological responses to far-red (FR) light. Analyses of the effect of nonphotosynthetic irradiation (low-intensity FR light) on petunia floral volatiles revealed FR light as a scent-activating factor. While PHYTOCHROME A regulated scent-related gene expression and floral scent production under FR light, the influence of PES on volatile production was not limited by FR light conditions.
© American Society of Plant Biologists 2023. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Conflict of interest statement
Conflict of interest statement. The authors declare no conflict of interest.
Figures
References
-
- Albo B. Transcription factor EOB1: a key component in a network regulating production of phenylpropanoid scent compounds in petunia flowers [thesis]. The Hebrew University of Jerusalem, Rehovot, Israel. 2012.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

