Multi-omics analysis of N6-methyladenosine reader IGF2BP3 as a promising biomarker in pan-cancer
- PMID: 36761737
- PMCID: PMC9905439
- DOI: 10.3389/fimmu.2023.1071675
Multi-omics analysis of N6-methyladenosine reader IGF2BP3 as a promising biomarker in pan-cancer
Abstract
Background: Insulin-like growth factor 2 mRNA-binding protein 3 (IGF2BP3) has been reported to exhibit an oncogenic effect as an RNA-binding protein (RBP) by promoting tumor cell proliferation, migration and invasion in several tumor types. However, a pan-cancer analysis of IGF2BP3 is not currently available, and the exact roles of IGF2BP3 in prognosis and immunology in cancer patients remain enigmatic. The main aim of this study was to provide visualization of the systemic prognostic landscape of IGF2BP3 in pan-cancer and to uncover the potential relationship between IGF2BP3 expression in the tumor microenvironment and immune infiltration profile.
Methods: Raw data on IGF2BP3 expression were obtained from GTEx, CCLE, TCGA, and HPA data portals. We have investigated the expression patterns, diagnostic and prognostic significance, mutation landscapes, functional analysis, and functional states of IGF2BP3 utilizing multiple databases, including HPA, TISIDB, cBioPortal, GeneMANIA, GESA, and CancerSEA. Moreover, the relationship of IGF2BP3 expression with immune infiltrates, TMB, MSI and immune-related genes was evaluated in pan-cancer. IGF2BP3 with drug sensitivity analysis was performed from the CellMiner database. Furthermore, the expression of IGF2BP3 in different grades of glioma was detected by immunohistochemical staining and western blot.
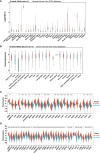
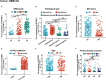
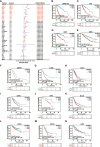
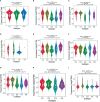
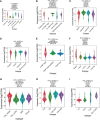
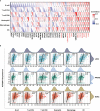
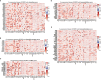
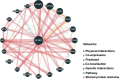
Results: We found that IGF2BP3 was ubiquitously highly expressed in pan-cancer and significantly correlated with diagnosis, prognosis, TMB, MSI, and drug sensitivity in various types of cancer. Besides, IGF2BP3 was involved in many cancer pathways and varied in different immune and molecular subtypes of cancers. Additionally, IGF2BP3 is critically associated with genetic markers of immunomodulators in various cancers. Finally, we validated that IGF2BP3 protein expression was significantly higher in glioma than in normal tissue, especially in GBM.
Conclusions: IGF2BP3 may be a potential molecular biomarker for diagnosis and prognosis in pan-cancer, especially for glioma. It could become a novel therapeutic target for various cancers.
Keywords: genetic alteration; immune infiltration; insulin-like growth factor 2 mRNA-binding protein 3 (IGF2BP3); pan-cancer analysis; prognosis; the Cancer Genome Atlas (TCGA).
Copyright © 2023 Chen, Xu, Cui, Wu, Xie and Zhang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures






Similar articles
-
Targeting IGF2BP3 in Cancer.Int J Mol Sci. 2023 May 29;24(11):9423. doi: 10.3390/ijms24119423. Int J Mol Sci. 2023. PMID: 37298373 Free PMC article. Review.
-
Multi-omics analysis of the oncogenic value of copper Metabolism-Related protein COMMD2 in human cancers.Cancer Med. 2023 May;12(10):11941-11959. doi: 10.1002/cam4.5320. Epub 2022 Oct 7. Cancer Med. 2023. PMID: 36205192 Free PMC article.
-
Pan-cancer evidence of prognosis, immune infiltration, and immunotherapy efficacy for annexin family using multi-omics data.Funct Integr Genomics. 2023 Jun 26;23(3):211. doi: 10.1007/s10142-023-01106-z. Funct Integr Genomics. 2023. PMID: 37358720
-
Comprehensive Pan-Cancer Genomic Analysis Reveals PHF19 as a Carcinogenic Indicator Related to Immune Infiltration and Prognosis of Hepatocellular Carcinoma.Front Immunol. 2022 Jan 5;12:781087. doi: 10.3389/fimmu.2021.781087. eCollection 2021. Front Immunol. 2022. PMID: 35069553 Free PMC article. Clinical Trial.
-
Cell Division Cycle-Associated Protein 3 (CDCA3) Is a Potential Biomarker for Clinical Prognosis and Immunotherapy in Pan-Cancer.Biomed Res Int. 2022 Aug 30;2022:4632453. doi: 10.1155/2022/4632453. eCollection 2022. Biomed Res Int. 2022. Retraction in: Biomed Res Int. 2023 Jun 21;2023:9870382. doi: 10.1155/2023/9870382. PMID: 36082153 Free PMC article. Retracted. Review.
Cited by
-
Identification of IGF2BP3 for the Expression and Prognosis in Gastrointestinal Cancers.Int J Med Sci. 2025 Jun 23;22(13):3191-3201. doi: 10.7150/ijms.114411. eCollection 2025. Int J Med Sci. 2025. PMID: 40765570 Free PMC article.
-
hnRNPH1: A Multifaceted Regulator in RNA Processing and Disease Pathogenesis.Int J Mol Sci. 2025 May 28;26(11):5159. doi: 10.3390/ijms26115159. Int J Mol Sci. 2025. PMID: 40507967 Free PMC article. Review.
-
A comprehensive prognostic and immune infiltration analysis of UBA1 in pan-cancer: A computational analysis and in vitro experiments.J Cell Mol Med. 2024 Aug;28(16):e70037. doi: 10.1111/jcmm.70037. J Cell Mol Med. 2024. PMID: 39183260 Free PMC article.
-
Targeting IGF2BP3 in Cancer.Int J Mol Sci. 2023 May 29;24(11):9423. doi: 10.3390/ijms24119423. Int J Mol Sci. 2023. PMID: 37298373 Free PMC article. Review.
-
Relationship between CTF1 gene expression and prognosis and tumor immune microenvironment in glioma.Eur J Med Res. 2025 Jan 9;30(1):17. doi: 10.1186/s40001-024-02192-w. Eur J Med Res. 2025. PMID: 39780198 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

