Expansion and loss of sperm nuclear basic protein genes in Drosophila correspond with genetic conflicts between sex chromosomes
- PMID: 36763410
- PMCID: PMC9917458
- DOI: 10.7554/eLife.85249
Expansion and loss of sperm nuclear basic protein genes in Drosophila correspond with genetic conflicts between sex chromosomes
Abstract
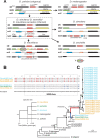
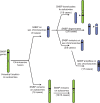
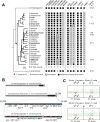
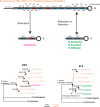
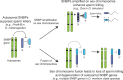
Many animal species employ sperm nuclear basic proteins (SNBPs) or protamines to package sperm genomes tightly. SNBPs vary across animal lineages and evolve rapidly in mammals. We used a phylogenomic approach to investigate SNBP diversification in Drosophila species. We found that most SNBP genes in Drosophila melanogaster evolve under positive selection except for genes essential for male fertility. Unexpectedly, evolutionarily young SNBP genes are more likely to be critical for fertility than ancient, conserved SNBP genes. For example, CG30056 is dispensable for male fertility despite being one of three SNBP genes universally retained in Drosophila species. We found 19 independent SNBP gene amplification events that occurred preferentially on sex chromosomes. Conversely, the montium group of Drosophila species lost otherwise-conserved SNBP genes, coincident with an X-Y chromosomal fusion. Furthermore, SNBP genes that became linked to sex chromosomes via chromosomal fusions were more likely to degenerate or relocate back to autosomes. We hypothesize that autosomal SNBP genes suppress meiotic drive, whereas sex-chromosomal SNBP expansions lead to meiotic drive. X-Y fusions in the montium group render autosomal SNBPs dispensable by making X-versus-Y meiotic drive obsolete or costly. Thus, genetic conflicts between sex chromosomes may drive SNBP rapid evolution during spermatogenesis in Drosophila species.
Keywords: D. melanogaster; evolutionary biology; genetics; genomics; meiotic drive; positive selection; protamines; pseudogenes.
Plain language summary
In sperm, DNA is packaged more tightly than in other cells thanks to small proteins called ‘sperm nuclear basic proteins’ (SNBPs), also called protamines in mammals. SNBPs are important for sperm to develop properly and correctly perform their role during fertilization. Although the evolution of SNBPs has been studied in mammals, these proteins have not been as thoroughly examined in invertebrates. Chang et al. took advantage of the availability of high-quality sequences for the genomes of 78 species of Drosophila flies to investigate the evolution of the genes that code for SNBPs in these flies. The results showed that, just like in mammals, in Drosophila the protein sequences of SNBPs evolve rapidly. However, unlike mammals, Chang et al. also found that Drosophila species frequently gained and lost genes coding for SNBPs. Interestingly, the ‘older’ genes (genes that appeared earlier in evolution) that code for SNBPs are not essential for reproduction in the fruit fly Drosophila melanogaster. This is an unexpected finding because older genes usually have essential roles for survival and reproduction, which require them to be passed on to the next generation and remain in the genome. In contrast, younger SNBP genes that had appeared more recently and were not shared between different species of Drosophila were often essential for fertility. These results, combined with other observations about where SNBP genes are located in the genome, led Chang et al. to hypothesize that SNBPs present in sex chromosomes act as ‘meiotic drivers’ while those on other chromosomes (known as autosomes) suppress meiotic drive. In other words, SNBP genes present in the sex chromosomes may be responsible for killing sister sperm cells that do not carry those genes, while SNBP genes that are not located on sex chromosomes may suppress this activity. This is of particular interest because it indicates that SNBPs are involved in genetic conflicts between the two sex chromosomes: sperm that carry SNBPs on the X chromosome may kill sperm with a Y chromosome, and vice versa. The results of Chang et al. shed light on the mysterious evolution of SNBPs in Drosophila flies. Although previous hypotheses regarding the rapid evolution of SNBPs evolution have focused on their role in genome packaging, this new analysis suggests that much of the evolutionary change is likely driven by genetic conflicts between sex chromosomes.
© 2023, Chang et al.
Conflict of interest statement
CC, IM, HM No competing interests declared
Figures

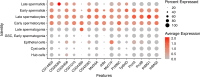
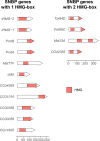



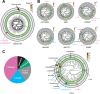
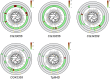

Similar articles
-
Sperm Nuclear Basic Proteins of Marine Invertebrates.Results Probl Cell Differ. 2018;65:15-32. doi: 10.1007/978-3-319-92486-1_2. Results Probl Cell Differ. 2018. PMID: 30083913
-
The Drosophila chromosomal protein Mst77F is processed to generate an essential component of mature sperm chromatin.Open Biol. 2016 Nov;6(11):160207. doi: 10.1098/rsob.160207. Open Biol. 2016. PMID: 27810970 Free PMC article.
-
The sperm nuclear basic proteins (SNBPs) of the sponge Neofibularia nolitangere: implications for the molecular evolution of SNBPs.J Mol Evol. 1997 Jul;45(1):91-6. doi: 10.1007/pl00006207. J Mol Evol. 1997. PMID: 9211739
-
Evolution of vertebrate chromosomal sperm proteins: implications for fertility and sperm competition.Soc Reprod Fertil Suppl. 2007;65:63-79. Soc Reprod Fertil Suppl. 2007. PMID: 17644955 Review.
-
Sex-ratio meiotic drive in Drosophila simulans: cellular mechanism, candidate genes and evolution.Biochem Soc Trans. 2006 Aug;34(Pt 4):562-5. doi: 10.1042/BST0340562. Biochem Soc Trans. 2006. PMID: 16856861 Review.
Cited by
-
Essential and recurrent roles for hairpin RNAs in silencing de novo sex chromosome conflict in Drosophila simulans.PLoS Biol. 2023 Jun 8;21(6):e3002136. doi: 10.1371/journal.pbio.3002136. eCollection 2023 Jun. PLoS Biol. 2023. PMID: 37289846 Free PMC article.
-
Comparative single-cell analysis of transcriptional bursting reveals the role of genome organization in de novo transcript origination.Proc Natl Acad Sci U S A. 2025 May 6;122(18):e2425618122. doi: 10.1073/pnas.2425618122. Epub 2025 Apr 30. Proc Natl Acad Sci U S A. 2025. PMID: 40305051
-
New perspectives on the causes and consequences of male meiotic drive.Curr Opin Genet Dev. 2023 Dec;83:102111. doi: 10.1016/j.gde.2023.102111. Epub 2023 Sep 11. Curr Opin Genet Dev. 2023. PMID: 37704518 Free PMC article. Review.
-
Trim66's paternal deficiency causes intrauterine overgrowth.Life Sci Alliance. 2024 May 7;7(7):e202302512. doi: 10.26508/lsa.202302512. Print 2024 Jul. Life Sci Alliance. 2024. PMID: 38719749 Free PMC article.
-
The roles of gene duplications in the dynamics of evolutionary conflicts.Proc Biol Sci. 2024 Jun;291(2024):20240555. doi: 10.1098/rspb.2024.0555. Epub 2024 Jun 12. Proc Biol Sci. 2024. PMID: 38865605 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

