A Comprehensive Study on Antibiotic Resistance among Coagulase-Negative Staphylococci (CoNS) Strains Isolated from Ready-to-Eat Food Served in Bars and Restaurants
- PMID: 36766043
- PMCID: PMC9914766
- DOI: 10.3390/foods12030514
A Comprehensive Study on Antibiotic Resistance among Coagulase-Negative Staphylococci (CoNS) Strains Isolated from Ready-to-Eat Food Served in Bars and Restaurants
Abstract
The present study aimed to characterize and assess the diversity of CoNS strains as potential vectors for the spread of resistance to antimicrobial agents from RTE foods served in bars and restaurants. Eighty-five CoNS strains, obtained from 198 RTE food samples, were investigated. Sixty-seven CoNS isolates (78.8%) were resistant to at least one antibiotic tested, and 37 (43.5%) were multidrug resistant (MDR-CoNS). Moreover, CoNS strains contained genes conferring resistance to antibiotics critically important in medicine, i.e., β-lactams [mecA (29.4%); blaZ (84.7%)], aminoglycosides [aac(6')-Ie-aph(2″)-Ia (45.9%); aph(2″)-Ic (3.5%)], macrolides, lincosamides and streptogramin B-MLSB [msrA/B (68.2%); ermB (40%) and mphC (4.7%)], tetracyclines [tetK (31.8%); tetM (16.5%) and/or tetL (2.35%)]. We also found the fusB/C/D genes responsible for the acquired low-level fusidic acid resistance (17.6%) and streptogramin resistance determinant vgaA in 30.6% of isolates. In three linezolid resistant strains (2 S. epidermidis and 1 S. warneri), mutation was detected, as demonstrated by L101V and V188I changes in the L3 protein amino acid sequences. The high frequency in RTE food of MDR-CoNS including methicillin-resistant (MR-CoNS) strains constitutes a direct risk to public health as they increase the gene pool from which pathogenic bacteria can pick up resistance traits.
Keywords: antimicrobial resistance; coagulase-negative staphylococci (CoNS); methicillin resistant coagulase-negative staphylococci (MR-CoNS); multidrug resistant coagulase-negative staphylococci (MDR-CoNS); ready-to-eat food.
Conflict of interest statement
The authors declare no conflict of interest.
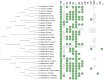
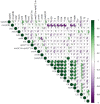
Figures
Similar articles
-
Occurrence and characteristics of methicillin-resistant and -susceptible Staphylococcus aureus and methicillin-resistant coagulase-negative staphylococci from Japanese retail ready-to-eat raw fish.Int J Food Microbiol. 2012 Jun 1;156(3):286-9. doi: 10.1016/j.ijfoodmicro.2012.03.022. Epub 2012 Mar 28. Int J Food Microbiol. 2012. PMID: 22541390
-
Antimicrobial resistome of coagulase-negative staphylococci from nasotracheal cavities of nestlings of Ciconia ciconia in Southern Spain: Detection of mecC-SCCmec type-XI-carrying S. lentus.Comp Immunol Microbiol Infect Dis. 2023 Aug;99:102012. doi: 10.1016/j.cimid.2023.102012. Epub 2023 Jun 30. Comp Immunol Microbiol Infect Dis. 2023. PMID: 37453201
-
Coagulase-negative staphylococci (CoNS) isolated from ready-to-eat food of animal origin--phenotypic and genotypic antibiotic resistance.Food Microbiol. 2015 Apr;46:222-226. doi: 10.1016/j.fm.2014.08.001. Epub 2014 Aug 23. Food Microbiol. 2015. PMID: 25475289
-
Phenotypic characteristics of coagulase-negative staphylococci: typing and antibiotic susceptibility.APMIS Suppl. 1999;91:1-42. APMIS Suppl. 1999. PMID: 10230367 Review.
-
Potentiation and Mechanism of Berberine as an Antibiotic Adjuvant Against Multidrug-Resistant Bacteria.Infect Drug Resist. 2023 Nov 21;16:7313-7326. doi: 10.2147/IDR.S431256. eCollection 2023. Infect Drug Resist. 2023. PMID: 38023403 Free PMC article. Review.
Cited by
-
Molecular epidemiology and characterization of antimicrobial-resistant Staphylococcus haemolyticus strains isolated from dairy cattle milk in Northwest, China.Front Cell Infect Microbiol. 2023 May 17;13:1183390. doi: 10.3389/fcimb.2023.1183390. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 37265496 Free PMC article.
-
Antibiotic Susceptibility Profiles of Bacterial Isolates Recovered from Abscesses in Cattle and Sheep at a Slaughterhouse in Algeria.Microorganisms. 2024 Mar 5;12(3):524. doi: 10.3390/microorganisms12030524. Microorganisms. 2024. PMID: 38543576 Free PMC article.
-
Identification of bacteria on Thai banknotes and coins using MALDI-TOF mass spectrometry and their phenotypic antimicrobial susceptibility profiles.PeerJ. 2025 May 20;13:e19465. doi: 10.7717/peerj.19465. eCollection 2025. PeerJ. 2025. PMID: 40416613 Free PMC article.
-
Characteristics of drug-resistant staphylococci isolated from milk of lambed ewes during the perinatal period.J Vet Res. 2025 Mar 25;69(1):41-50. doi: 10.2478/jvetres-2025-0014. eCollection 2025 Mar. J Vet Res. 2025. PMID: 40144065 Free PMC article.
-
Prevalence and Antibiotic Resistance of Bacillus sp. Isolated from Raw Milk.Microorganisms. 2023 Apr 19;11(4):1065. doi: 10.3390/microorganisms11041065. Microorganisms. 2023. PMID: 37110488 Free PMC article.
References
-
- Pedroso S.H., Sandes S.H., Filho R.A., Nunes A.C., Serufo J.C., Farias L.M., Carvalho A.R., Bomfim R.Q.M., Santos S.G. Coagulase-negative Staphylococci isolated from human bloodstream infections showed multidrug resistance profile. Microb. Drug Resist. 2018;24:5–635. doi: 10.1089/mdr.2017.0309. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

