Molecular Toxicity Virtual Screening Applying a Quantized Computational SNN-Based Framework
- PMID: 36771009
- PMCID: PMC9919191
- DOI: 10.3390/molecules28031342
Molecular Toxicity Virtual Screening Applying a Quantized Computational SNN-Based Framework
Abstract
Spiking neural networks are biologically inspired machine learning algorithms attracting researchers' attention for their applicability to alternative energy-efficient hardware other than traditional computers. In the current work, spiking neural networks have been tested in a quantitative structure-activity analysis targeting the toxicity of molecules. Multiple public-domain databases of compounds have been evaluated with spiking neural networks, achieving accuracies compatible with high-quality frameworks presented in the previous literature. The numerical experiments also included an analysis of hyperparameters and tested the spiking neural networks on molecular fingerprints of different lengths. Proposing alternatives to traditional software and hardware for time- and resource-consuming tasks, such as those found in chemoinformatics, may open the door to new research and improvements in the field.
Keywords: in silico toxicity prediction; machine learning; molecular fingerprints; spiking neural networks.
Conflict of interest statement
The authors declare no conflict of interest.
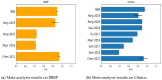
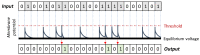
Figures





Similar articles
-
Compact Hardware Synthesis of Stochastic Spiking Neural Networks.Int J Neural Syst. 2019 Oct;29(8):1950004. doi: 10.1142/S0129065719500047. Epub 2019 Feb 8. Int J Neural Syst. 2019. PMID: 30880526
-
Advancements in Algorithms and Neuromorphic Hardware for Spiking Neural Networks.Neural Comput. 2022 May 19;34(6):1289-1328. doi: 10.1162/neco_a_01499. Neural Comput. 2022. PMID: 35534005 Review.
-
Supervised learning in spiking neural networks: A review of algorithms and evaluations.Neural Netw. 2020 May;125:258-280. doi: 10.1016/j.neunet.2020.02.011. Epub 2020 Feb 25. Neural Netw. 2020. PMID: 32146356 Review.
-
Overview of Spiking Neural Network Learning Approaches and Their Computational Complexities.Sensors (Basel). 2023 Mar 11;23(6):3037. doi: 10.3390/s23063037. Sensors (Basel). 2023. PMID: 36991750 Free PMC article. Review.
-
BindsNET: A Machine Learning-Oriented Spiking Neural Networks Library in Python.Front Neuroinform. 2018 Dec 12;12:89. doi: 10.3389/fninf.2018.00089. eCollection 2018. Front Neuroinform. 2018. PMID: 30631269 Free PMC article.
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

