Three-Dimensional-QSAR and Relative Binding Affinity Estimation of Focal Adhesion Kinase Inhibitors
- PMID: 36771129
- PMCID: PMC9919860
- DOI: 10.3390/molecules28031464
Three-Dimensional-QSAR and Relative Binding Affinity Estimation of Focal Adhesion Kinase Inhibitors
Abstract
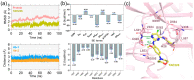
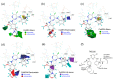
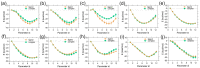
Precise binding affinity predictions are essential for structure-based drug discovery (SBDD). Focal adhesion kinase (FAK) is a member of the tyrosine kinase protein family and is overexpressed in a variety of human malignancies. Inhibition of FAK using small molecules is a promising therapeutic option for several types of cancer. Here, we conducted computational modeling of FAK-targeting inhibitors using three-dimensional structure-activity relationship (3D-QSAR), molecular dynamics (MD), and hybrid topology-based free energy perturbation (FEP) methods. The structure-activity relationship (SAR) studies between the physicochemical descriptors and inhibitory activities of the chemical compounds were performed with reasonable statistical accuracy using CoMFA and CoMSIA. These are two well-known 3D-QSAR methods based on the principle of supervised machine learning (ML). Essential information regarding residue-specific binding interactions was determined using MD and MM-PB/GBSA methods. Finally, physics-based relative binding free energy (ΔΔGRBFEA→B) terms of analogous ligands were estimated using alchemical FEP simulation. An acceptable agreement was observed between the experimental and computed relative binding free energies. Overall, the results suggested that using ML and physics-based hybrid approaches could be useful in synergy for the rational optimization of accessible lead compounds with similar scaffolds targeting the FAK receptor.
Keywords: 3D-QSAR; MM-PB/GBSA; focal adhesion kinase; free energy perturbation; molecular dynamics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Identification of potential FAK inhibitors using mol2vec molecular descriptor-based QSAR, molecular docking, ADMET study, and molecular dynamics simulation.Mol Divers. 2024 Aug;28(4):2163-2175. doi: 10.1007/s11030-024-10839-3. Epub 2024 Apr 6. Mol Divers. 2024. PMID: 38582821
-
In silico design of novel FAK inhibitors using integrated molecular docking, 3D-QSAR and molecular dynamics simulation studies.J Biomol Struct Dyn. 2022 Aug;40(13):5965-5982. doi: 10.1080/07391102.2021.1875880. Epub 2021 Jan 21. J Biomol Struct Dyn. 2022. PMID: 33475043
-
Binding Studies and Lead Generation of Pteridin-7(8H)-one Derivatives Targeting FLT3.Int J Mol Sci. 2022 Jul 12;23(14):7696. doi: 10.3390/ijms23147696. Int J Mol Sci. 2022. PMID: 35887060 Free PMC article.
-
Investigations of FAK inhibitors: a combination of 3D-QSAR, docking, and molecular dynamics simulations studies.J Biomol Struct Dyn. 2018 May;36(6):1529-1549. doi: 10.1080/07391102.2017.1329095. Epub 2017 May 31. J Biomol Struct Dyn. 2018. PMID: 28490269
-
Tracking protein kinase targeting advances: integrating QSAR into machine learning for kinase-targeted drug discovery.Future Sci OA. 2025 Dec;11(1):2483631. doi: 10.1080/20565623.2025.2483631. Epub 2025 Apr 4. Future Sci OA. 2025. PMID: 40181786 Free PMC article. Review.
Cited by
-
Aurora kinase A promotes epithelial‑mesenchymal transition by regulating P130 and P107 molecules in thyroid cancer cells.Exp Ther Med. 2025 Mar 12;29(5):93. doi: 10.3892/etm.2025.12843. eCollection 2025 May. Exp Ther Med. 2025. PMID: 40162054 Free PMC article.
-
New strategies to enhance the efficiency and precision of drug discovery.Front Pharmacol. 2025 Feb 11;16:1550158. doi: 10.3389/fphar.2025.1550158. eCollection 2025. Front Pharmacol. 2025. PMID: 40008135 Free PMC article. Review.
-
Identification of potential FAK inhibitors using mol2vec molecular descriptor-based QSAR, molecular docking, ADMET study, and molecular dynamics simulation.Mol Divers. 2024 Aug;28(4):2163-2175. doi: 10.1007/s11030-024-10839-3. Epub 2024 Apr 6. Mol Divers. 2024. PMID: 38582821
References
-
- Liao Y., Liu L., Yang J., Shi Z. ATX/LPA axis regulates FAK activation, cell proliferation, apoptosis, and motility in human pancreatic cancer cells. In Vitro Cell. Dev. Biol. Anim. 2022;58:307–315. - PubMed
-
- Le Coq J., Acebrón I., Martin B.R., Navajas P.L., Lietha D. New insights into FAK structure and function in focal adhesions. J. Cell Sci. 2022;135:jcs259089. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

