Genome-Wide Association Study of Rice Grain Shape and Chalkiness in a Worldwide Collection of Xian Accessions
- PMID: 36771503
- PMCID: PMC9919668
- DOI: 10.3390/plants12030419
Genome-Wide Association Study of Rice Grain Shape and Chalkiness in a Worldwide Collection of Xian Accessions
Abstract
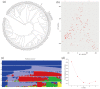
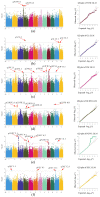
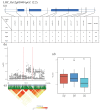
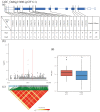
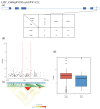
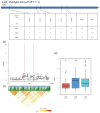
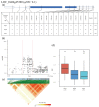
Rice (Oryza sativa L.) appearance quality, which is mainly defined by grain shape and chalkiness, is an important target in rice breeding. In this study, we first re-sequenced 137 indica accessions and then conducted a genome-wide association study (GWAS) for six agronomic traits with the 2,998,034 derived single nucleotide polymorphisms (SNPs) by using the best linear unbiased prediction (BLUP) values for each trait. The results revealed that 195 SNPs had significant associations with the six agronomic traits. Based on the genome-wide linkage disequilibrium (LD) blocks, candidate genes for the target traits were detected within 100 kb upstream and downstream of the relevant SNP loci. Results indicate that six quantitative trait loci (QTLs) significantly associated with six traits (qTGW4.1, qTGW4.2, qGL4.1, qGL12.1, qGL12.2, qGW2.1, qGW4.1, qGW6.1, qGW8.1, qGW8.2, qGW9.1, qGW11.1, qGLWR2.1, qGLWR2.2, qGLWR4.2, qPGWC5.1 and qDEC6.1) were identified for haplotype analysis. Among these QTLs, two (qTGW4.2 and qGW6.1), were overlapped with FLO19 and OsbZIP47, respectively, and the remaining four were novel QTLs. These candidate genes were further validated by haplotype block construction.
Keywords: GWAS; QTLs; chalkiness; grain shape; rice.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Fao R.I.E. Increasing Crop Production Sustainably: The Perspective of Biological of Processes. Food and Agriculture Organization; Rome, Italy: 2009.
-
- Godfray H.C.J. The challenge of feeding 9-10 billion people equitably and sustainably. J. Agric. Sci. 2014;152:S2–S8. doi: 10.1017/S0021859613000774. - DOI
-
- Seck P.A., Diagne A., Mohanty S., Wopereis M. Crops that feed the world 7: Rice. Food Secur. 2012;4:7–24. doi: 10.1007/s12571-012-0168-1. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

