The ubiquitination landscape of the influenza A virus polymerase
- PMID: 36774438
- PMCID: PMC9922279
- DOI: 10.1038/s41467-023-36389-0
The ubiquitination landscape of the influenza A virus polymerase
Abstract
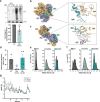
During influenza A virus (IAV) infections, viral proteins are targeted by cellular E3 ligases for modification with ubiquitin. Here, we decipher and functionally explore the ubiquitination landscape of the IAV polymerase proteins during infection of human alveolar epithelial cells by applying mass spectrometry analysis of immuno-purified K-ε-GG (di-glycyl)-remnant-bearing peptides. We have identified 59 modified lysines across the three subunits, PB2, PB1 and PA of the viral polymerase of which 17 distinctively affect mRNA transcription, vRNA replication and the generation of recombinant viruses via non-proteolytic mechanisms. Moreover, further functional and in silico analysis indicate that ubiquitination at K578 in the PB1 thumb domain is mechanistically linked to dynamic structural transitions of the viral polymerase that are required for vRNA replication. Mutations K578A and K578R differentially affect the generation of recombinant viruses by impeding cRNA and vRNA synthesis, NP binding as well as polymerase dimerization. Collectively, our results demonstrate that the ubiquitin-mediated charge neutralization at PB1-K578 disrupts the interaction to an unstructured loop in the PB2 N-terminus that is required to coordinate polymerase dimerization and facilitate vRNA replication. This provides evidence that IAV exploits the cellular ubiquitin system to modulate the activity of the viral polymerase for viral replication.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





References
-
- Ruigrok RW, Baudin F. Structure of influenza virus ribonucleoprotein particles. II. Purified RNA-free influenza virus ribonucleoprotein forms structures that are indistinguishable from the intact influenza virus ribonucleoprotein particles. J. Gen. Virol. 1995;76:1009–1014. doi: 10.1099/0022-1317-76-4-1009. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

