This is a preprint.
Population analyses of mosaic X chromosome loss identify genetic drivers and widespread signatures of cellular selection
- PMID: 36778285
- PMCID: PMC9915812
- DOI: 10.1101/2023.01.28.23285140
Population analyses of mosaic X chromosome loss identify genetic drivers and widespread signatures of cellular selection
Abstract
Mosaic loss of the X chromosome (mLOX) is the most commonly occurring clonal somatic alteration detected in the leukocytes of women, yet little is known about its genetic determinants or phenotypic consequences. To address this, we estimated mLOX in >900,000 women across eight biobanks, identifying 10% of women with detectable X loss in approximately 2% of their leukocytes. Out of 1,253 diseases examined, women with mLOX had an elevated risk of myeloid and lymphoid leukemias and pneumonia. Genetic analyses identified 49 common variants influencing mLOX, implicating genes with established roles in chromosomal missegregation, cancer predisposition, and autoimmune diseases. Complementary exome-sequence analyses identified rare missense variants in FBXO10 which confer a two-fold increased risk of mLOX. A small fraction of these associations were shared with mosaic Y chromosome loss in men, suggesting different biological processes drive the formation and clonal expansion of sex chromosome missegregation events. Allelic shift analyses identified alleles on the X chromosome which are preferentially retained, demonstrating that variation at many loci across the X chromosome is under cellular selection. A novel polygenic score including 44 independent X chromosome allelic shift loci correctly inferred the retained X chromosomes in 80.7% of mLOX cases in the top decile. Collectively our results support a model where germline variants predispose women to acquiring mLOX, with the allelic content of the X chromosome possibly shaping the magnitude of subsequent clonal expansion.
Figures
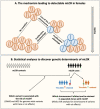



References
-
- Brown C.J., Ballabio A., Rupert J.L., Lafreniere R.G., Grompe M., Tonlorenzi R. and Willard H.F., 1991. A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome. Nature, 349(6304), pp.38–44. - PubMed
-
- Lyon M.F., 1961. Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature, 190(4773), pp.372–373. - PubMed
-
- Busque L., Mio R., Mattioli J., Brais E., Blais N., Lalonde Y., Maragh M. and Gilliland D.G., 1996. Nonrandom X-inactivation patterns in normal females: lyonization ratios vary with age. Blood, 88, 59–65. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
