Development and application of a simple pharmacokinetic model that quantitatively describes the distribution and elimination of the commonly measured proteins
- PMID: 36778906
- PMCID: PMC9909726
- DOI: 10.5599/admet.1570
Development and application of a simple pharmacokinetic model that quantitatively describes the distribution and elimination of the commonly measured proteins
Abstract
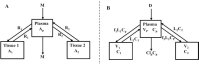
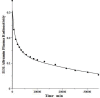
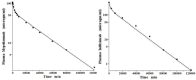
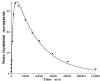
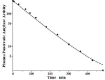
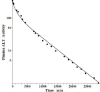
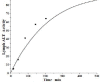
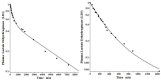
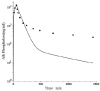
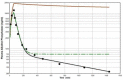
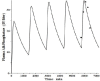
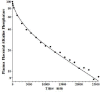
Increased plasma concentrations of a variety of cellular enzymes (alanine transaminase, aspartate aminotransferase, alkaline phosphatase, amylase, etc.) are commonly used as routine screening tests for a range of conditions. An increased concentration usually is assumed to result from an increased rate of delivery to the plasma. Factors such as decreased metabolism or excretion or altered extravascular distribution usually are ignored. As a prelude to a detailed analysis of all the factors producing altered plasma enzyme levels, we have reviewed the relevant literature describing the pharmacokinetics (PK) of 13 of the commonly measured plasma proteins and developed a PK model that provides a simple physiological description of all the data. Our model starts with the general 3-compartment, 6-parameter system previously developed for albumin and interprets the fluxes in terms of unidirectional sieved protein convectional volume flows from the plasma to the two tissue compartments and equal lymph flows returning to the plasma. This greatly constrains the model such that each protein is characterized by only two adjustable parameters (plasma clearance and sieving factor). In addition to accurately fitting the plasma kinetics, the model can accurately describe the tissue and lymph protein PK. For example, it can describe the thoracic duct lymph protein concentration following an intravenous infusion or the plasma concentration following a subcutaneous tissue injection. This simple model provides a satisfactory framework for the PK of 12 of the 13 proteins investigated. The glycoprotein intestinal alkaline phosphatase is the exception, requiring the addition of a liver recycling compartment involving the asialoglycoprotein receptor.
Keywords: Protein; alanine transaminase; albumin; alkaline phosphatase; amylase; aspartate aminotransferase; liver; pancreas; pharmacokinetics.
Copyright © 2023 by the authors.
Conflict of interest statement
Conflict of interest : The authors report no conflicts of interest in this work.
Figures


Similar articles
-
Comparing models for perfluorooctanoic acid pharmacokinetics using Bayesian analysis.J Pharmacokinet Pharmacodyn. 2008 Dec;35(6):683-712. doi: 10.1007/s10928-008-9108-2. Epub 2009 Jan 8. J Pharmacokinet Pharmacodyn. 2008. PMID: 19130186
-
Isoniazid plasma and lymph kinetics after intravenous injection in dogs.Arzneimittelforschung. 1984;34(10):1283-8. Arzneimittelforschung. 1984. PMID: 6542789
-
Population pharmacokinetics meta-analysis of plitidepsin (Aplidin) in cancer subjects.Cancer Chemother Pharmacol. 2009 Jun;64(1):97-108. doi: 10.1007/s00280-008-0841-4. Epub 2008 Oct 22. Cancer Chemother Pharmacol. 2009. PMID: 18941750
-
[Compartment simulation models and physiologic use in pharmaco- and toxicokinetics].Med Pr. 1998;49(1):83-92. Med Pr. 1998. PMID: 9587914 Review. Polish.
-
[Modern concepts in pharmacokinetics of intravenous anesthetics].Anaesthesist. 2004 Jan;53(1):95-110. doi: 10.1007/s00101-003-0628-2. Anaesthesist. 2004. PMID: 14994741 Review. German.
Cited by
-
Novel extracorporeal treatment for severe neonatal jaundice: a mathematical modelling study of allo-hemodialysis.Sci Rep. 2024 Sep 19;14(1):21910. doi: 10.1038/s41598-024-72256-8. Sci Rep. 2024. PMID: 39300159 Free PMC article.
References
-
- Brandse J.F., Mould D., Smeekes O., Ashruf Y., Kuin S., Strik A., van den Brink G.R., D'Haens G.R.. A Real-life Population Pharmacokinetic Study Reveals Factors Associated with Clearance and Immunogenicity of Infliximab in Inflammatory Bowel Disease. Inflamm Bowel Dis 23 (2017) 650-660. https://doi.org/10.1097/MIB.0000000000001043 10.1097/MIB.0000000000001043. - DOI - PubMed
-
- Fasanmade A.A., Adedokun O.J., Ford J., Hernandez D., Johanns J., Hu C., Davis H.M., Zhou H.. Population pharmacokinetic analysis of infliximab in patients with ulcerative colitis. Eur J Clin Pharmacol 65 (2009) 1211-1228. https://doi.org/10.1007/s00228-009-0718-4 10.1007/s00228-009-0718-4. - DOI - PMC - PubMed
-
- Beeken W.L., Volwiler W., Goldsworthy P.D., Garby L.E., Reynolds W.E., Stogsdill R., Stemler R.S.. Studies of I-131-albumin catabolism and distribution in normal young male adults. J Clin Invest 41 (1962) 1312-1333. https://doi.org/10.1172/JCI104594 10.1172/JCI104594. - DOI - PMC - PubMed
-
- Takeda Y., Reeve E.B.. Studies of the metabolism and distribution of albumin with autologous I131-albumin in healthy men. J Lab Clin Med 61 (1963) 183-202. https://www.ncbi.nlm.nih.gov/13984638 . - PubMed
-
- Rose R.H., Sepp A., Stader F., Gill K.L., Liu C., Gardner I.. Application of physiologically-based pharmacokinetic models for therapeutic proteins and other novel modalities. Xenobiotica (2022) 1-27. https://doi.org/10.1080/00498254.2022.2133649 10.1080/00498254.2022.2133649. - DOI - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
