Rare-variant association analysis reveals known and new age-related hearing loss genes
- PMID: 36788145
- PMCID: PMC10250305
- DOI: 10.1038/s41431-023-01302-2
Rare-variant association analysis reveals known and new age-related hearing loss genes
Abstract
Age-related (AR) hearing loss (HL) is a prevalent sensory deficit in the elderly population. Several studies showed that common variants increase ARHL susceptibility. Here, we demonstrate that rare-variants play a crucial role in ARHL etiology. We analyzed exome and imputed data from white-European UK Biobank volunteers, performing both single-variant and rare-variant aggregate association analyses using self-reported ARHL phenotypes. We identified and replicated associations between ARHL and rare-variants in KLHDC7B, PDCD6, MYO6, SYNJ2, and TECTA. PUS7L and EYA4 also revealed rare-variant associations with ARHL. EYA4, MYO6, and TECTA are all known to underline Mendelian nonsyndromic HL. PDCD6, a new HL gene, plays an important role in apoptosis and has widespread inner ear expression, particularly in the inner hair cells. An unreplicated common variant association was previously observed for KHLDC7B, here we demonstrate that rare-variants in this gene also play a role in ARHL etiology. Additionally, the first replicated association between SYNJ2 and ARHL was detected. Analysis of common variants revealed several previously reported, i.e., ARHGEF28, and new, i.e., PIK3R3, ARHL associations, as well as ones we replicate here for the first time, i.e., BAIAP2L2, CRIP3, KLHDC7B, MAST2, and SLC22A7. It was also observed that the odds ratios for rare-variant ARHL associations, were higher than those for common variants. In conclusion, we demonstrate the vital role rare-variants, including those in Mendelian nonsyndromic HL genes, play in the etiology of ARHL.
© 2023. The Author(s), under exclusive licence to European Society of Human Genetics.
Conflict of interest statement
The authors declare no competing interests.
Figures
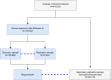


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

